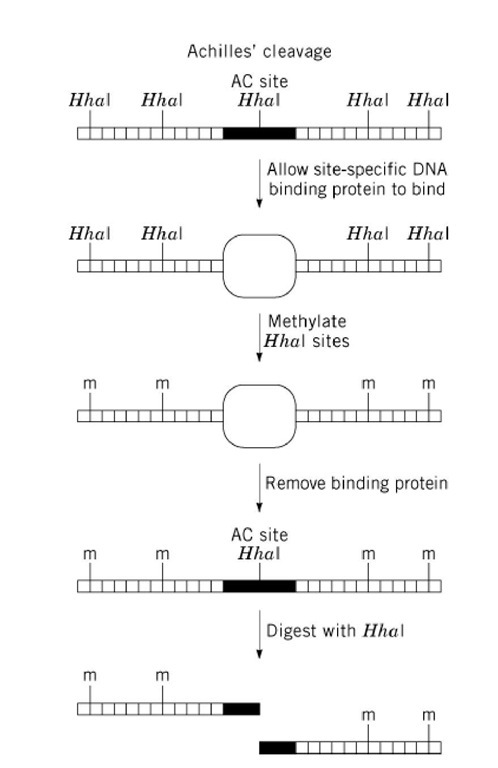
Achilles’ cleavage (AC) is a procedure developed to permit cutting double-stranded DNA at a single site (the AC site) in a complex genome (Fig. 1). The AC site is first protected from methylation by a sequence-specific DNA-binding protein, and the methylated genome is subsequently cut by a methylation-sensitive restriction enzyme. Other sites in the genome containing the same restriction/methylation site as the AC site are methylated and thus are not cut. The two key components of any AC system are (i) a restriction site that is recognized by both a methylation-sensitive restriction enzyme and its corresponding methyltransferase and (ii) the ability to block methylation of the chosen AC site by sequence-specific binding of a protein to that site. The overlap of the AC site and the recognition sequence for the DNA binding protein can occur naturally, or it can be engineered using recombination or site-directed mutagenesis. These requirements somewhat limit the applications of the AC technique.
Figure 1. Achilles’ Cleavage. Diagram of the method used to prevent restriction enzyme digestion at all sites except for the Achilles’ cleavage (AC) site. Horizontal lines connected by short vertical lines represent double-stranded DNA. HhaI restriction sites are marked by bold vertical lines above the double-stranded DNA. The binding site for the DNA-binding protein is represented by a solid rectangle. The DNA-binding protein is represented by an empty oval. Sites of methylation are indicated by a vertical line and the letter ‘m.’
Koob et al. (1) originally developed this procedure using two different repressor/operator systems. In one instance, plasmids carrying the operator of the lac operon were incubated with the Lac repressor and then methylated by the Hha I methyltransferase. Repressor binding prevented methylation of the Hha I site located within the lac operator. The methyltransferase and the operator protein were subsequently removed, leaving only the unmethylated AC site free to be cut by Hha I (1). Variations of the original AC method were also used to demonstrate the ability to make only one or a few cuts in the genomes of yeast and Escherichia coli (2), to turn frequent-cutting restriction enzymes into rare cutters (3), and to analyze the strength of protein-DNA interactions (4).
1. RecA-AC and RARE
The original AC method was made more universally applicable by using RecA protein and a sequence-specific oligonucleotide to form the protective complex, obviating the need for a sequence-specific binding protein. Two groups simultaneously developed this procedure, and their protocols were designated RecA-AC (5) and RARE (6). The RecA-AC/RARE technique still requires selection of a cleavage site that is recognized by both a restriction enzyme and its cognate methyltransferase. An oligonucleotide of 40 to 60 nucleotides in length is designed that matches the DNA sequence containing the cleavage site. In the presence of Mg and a non-hydrolyzable ATP analogue [eg, ATP(g-S)], RecA protein will bind to the oligonucleotide, forming a complex. When double-stranded DNA is added, the complex will bind to the matching genomic sequence to form a so-called ‘triple-stranded’ synaptic complex. This complex protects the cleavage site from methylation. As in the AC procedure, the methyltransferase and the RecA protein are removed after methylation, before the methylation-protected site is cleaved with the appropriate restriction enzyme.
As an alternative, the RecA complex can be used to protect specific restriction sites from restriction enzyme digestion, rather than from methyltransferase. This technique can be useful in some cloning procedures.
The RecA-AC/RARE technique has been used in various genome mapping experiments, for example to map gaps in human contiguous (contig) DNA and to map telomeres (7). The ability to cut genomic DNA at a few specific sites allows mapping large pieces of DNA by using pulsed-field gel electrophoresis and Southern blotting techniques to size DNA between two cleavage sites, or between a marker and the end of a telomere (7).
The RecA-AC/RARE technique is a useful technique for mapping and manipulating DNA and should continue to prove helpful in large-scale genome analysis studies of the future. One limitation to the technique is the number of sequences recognized by both a restriction enzyme and a methyltransferase. As more restriction enzyme/methyltransferase pairs are isolated or developed, the technique will be come even more useful.