Electrophoresis of fully denatured and dissociated polypeptide chains derived from proteins by binding of, and saturation with, the negatively charged detergent SDS, usually designated SDS-PAGE, is by far the most popular mode of gel electrophoresis in polyacrylamide gels (usually abbreviated PAGE). There are two reasons why SDS-PAGE is preferred: (1) it provides the resolving capacity and molecular sieving of PAGE, especially when conducted as disc electrophoresis (Fig. 1); and (2) it can give an estimate of the molecular weight of each polypeptide chain on the basis of electrophoresis at a single gel concentration. Molecular weights are often measured for protein using this technique alone, but it must be remembered that the results are valid only to the extent that the following simplifying conditions apply:
1. All SDS-denatured proteins tend to have the same net negative charge density: The binding of SDS to the protein usually masks the intrinsic charge of the protein and because of the bound SDS, gives it a standard negative charge density, that is largely independent of pH.
2. All SDS-denatured proteins tend to have the same conformation: The binding of SDS, when coupled with reduction of any disulfide bonds and dissociation into individual polypeptide chains at high temperatures, produces a conformation like a random coil for all SDS-polypeptides; therefore, mobility differences due to specific protein conformations are abolished.
3. The mobility is inversely proportional to the size of the polypeptide chain: Because the surface charge densities of proteins are equalized [see (1)], the Ferguson plots of all SDS-polypeptides should intersect at zero gel concentration; in this case, the ratios of the mobilities for all proteins at various gel concentrations are the same.
4. The mobility is inversely proportional to the logarithm of the molecular weight: A plot of log (molecular weight) versus migration distance is linear except at very large and very small molecular weights. Such a linear plot provides a simple means of deriving the molecular weight of an unknown protein (such as rhodopsin in the example of Fig. 2) by comparison to a set of standard proteins.
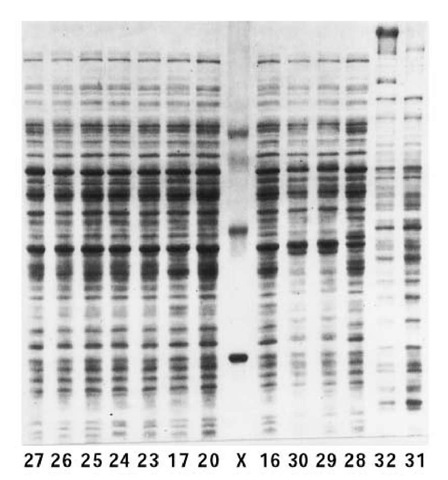
Figure 1. SDS-PAGE patterns of polypeptides derived from total cell extracts of various strains (numbered) of Achromobacter. Lane X contains the protein standards (from top to bottom) ovotransferrin (77 kDa), albumin (66 kDa), ovalbumin (43 kDa), carbonic anhydrase (30 kDa), and myoglobin (17 kDa). The polyacrylamide gel was of 10% (w/v) acrylamide, 2.7% Bisacrylamide, in a discontinuous Tris-chloride-glycinate buffer system.
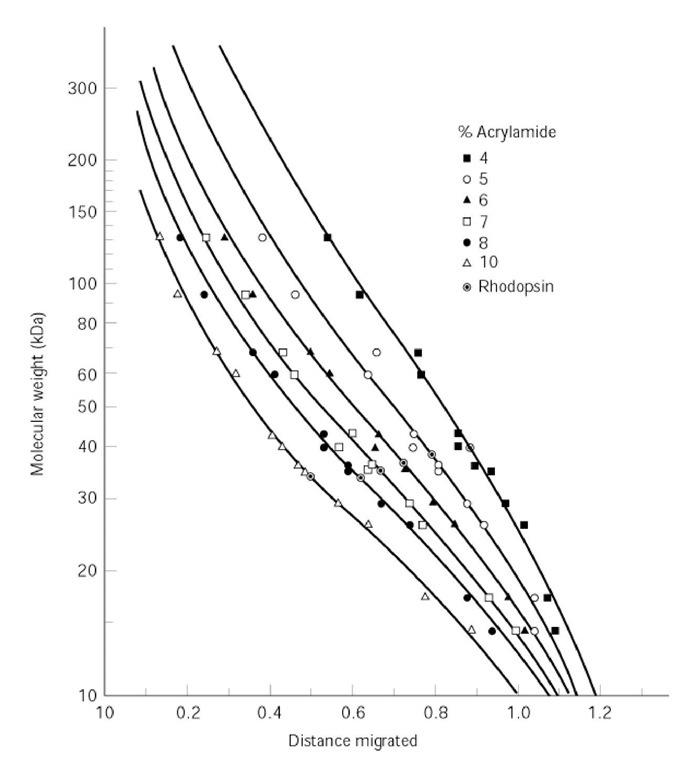
Figure 2. Standard curve of log(molecular weight) versus relative distance migrated in SDS-PAGE. Twelve standard proteins of known size were separated by SDS-PAGE in polyacrylamide gels of six different concentrations. The protein of molecular weight to be determined was rhodopsin.
The four simplifying conditions of SDS-PAGE given above are, however, not entirely or always valid, since:
1. The mobilities are usually constant between pH 7 and 11 but not at lower pH values.
2. The derivatization of proteins with SDS produces polypeptides of uniform random-coiled conformations only when the reaction conditions are sufficiently severe. In particular, the reaction time needs to be sufficient, which in many cases requires prolonged boiling with SDS. Once conformational uniformity and an approximately uniform net charge are attained (see condition 3 above), the mobilities and separations are more reproducible than those of native proteins.
3. Equalization of the surface charge densities of polypeptides by reaction with SDS is incomplete for proteins with extreme net charges or for those containing glyco-, lipo-, or nucleo-moieties. Such incomplete saturation of the protein surface with SDS accounts for the failure of the Ferguson plots of many SDS-polypeptides to intersect at or near the ordinate (1).
4. When the charge densities among proteins are not fully equalized by derivatization with SDS, the ratio of mobilities between size standards and unknowns becomes dependent on gel concentration. Therefore, the molecular weight obtained from a comparison with the mobilities of standards will vary with gel concentration; in that case, accurate molecular weights of SDS-polypeptides, like those of native proteins, can be derived only from the slope of a Ferguson plot (2).
5. Even if surface charge and conformational equalization among proteins are established for particular proteins, the sigmoidal nature of the plot of log(molecular weight) versus migration distance (Fig. 2) needs to be recognized (2). Thus, the approximation of linearity in the central segment of the sigmoidal curve needs to be restricted to a relatively narrow range of migration distances.
There are other common problems with SDS-PAGE:
1. Its SDS concentrations are greater than the critical micelle concentration (cmc), so that the SDS is present as micelles. Micellar SDS bound to the tracking dye unstacks at high gel concentrations in disc electrophoresis; so a tracking dye that marks the moving boundary front at low gel concentrations fails to do so at high concentrations (1).
2. Polyacrylamide is oxidative, and so disulfide bonds may be re-formed after proteins enter the gel; also, oxidation of agents used to reduce any disulfide bonds in the original sample may introduce artifactual new components in the gel pattern.
3. There is no theoretical understanding of SDS-PAGE in discontinuous buffer systems that takes account of moving boundaries set up by monomeric or micellar SDS and its complexes with buffer constituents.
For the foregoing reasons, it is always advisable to establish the accuracy of any important molecular weight value by Ferguson plot analysis.