FAMILY BORNAVIRIDAE
occurs to form an incompletely characterized set of alter-
natively spliced mRNAs from the five genes. Furthermore,
alternative start codons are used during translation of some
Borna disease virus is the sole representative of the fam-
of the mRNAs, a trait shared with other (-)RNA viruses, so
ily Bornaviridae in the order Mononegavirales (Table 4.6).
that overlapping open reading frames are present in some of
It has a nonsegmented, minus-sense genome of 8.9 kb, con-
the genes. Overall, the number of protein products produced
taining, by one definition, five genes. The general organi-
and the complexity of the readout of the genome exceed that
zation of the genome and the suite of genes contained
of other (-)RNA viruses. The genome organization and tran-
in the genome resembles that of other members of the
scriptional map of Borna disease virus, as currently under-
Mononegavirales (Fig. 4.1). However, the virus replicates
stood, is shown in Fig. 4.12.
in the nucleus, not in the cytoplasm. Splicing of mRNAs
TABLE 4.6 Bornaviridae
Virus name
World
Usual host(s)a
Genus/members
abbreviation
Transmission
Disease
distribution
Bornavirus
Borna disease
BDV
Horses, sheep other
??
Encephalopathy,
Europe, possibly
mammals
fatal paralysis
worldwide
a
It was recently reported that the bicolored white-toothed shrew Crocidura leucodon is the reservoir host in Switzerland where the disease is endemic. These
animals live on the ground and do not climb. Presumably horses are infected when grazing on forage contaminated by excretions from infected shrews;
horses fed exclusively from feeding troughs in modern barns are seldom affected (Hilbe et al., 2006 ).
Frame 3
N (p40/p38)
P (p24)
M(p16)
L (p180/p190)
Frame 2
ORFs
(p10)
Frame 1
Gn(gp50) Gc(gp43)
1
8908
39 OH
59
Genome
RNA
kb
0
1
2
3
4
5
6
7
8
9
N(p40)
mRNAs
N(p38)
X
P(p24)
P9
M(p16)
I
Gn
Gc
II
M(p16)
M(p16)
I
Gn
Gc
I
II
L (p190)
L (p180)
Transcriptional start site
Transcriptional end site
Translation initation
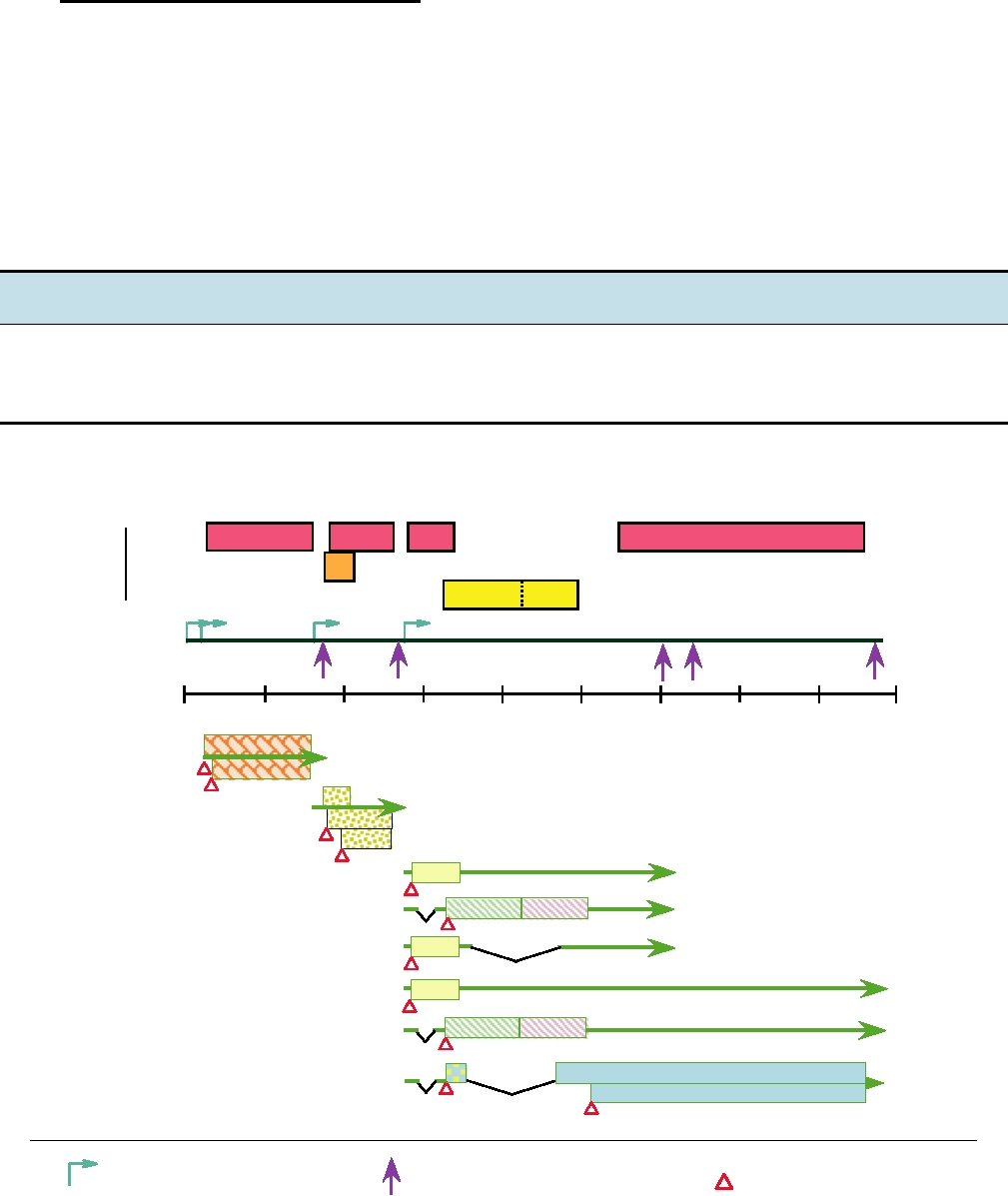
FIGURE 4.12 Genome organization and transcriptional map of an isolate of Borna disease virus. ORFs are represented
by the boxes at the top, color coded by reading frame. Messenger RNAs are shown below, with the boxes color coded
according to the function of the product as in Figs. 4.1 and 4.6. The positions of two introns, I (nucleotides 1932 to 2025)
and II (nucleotides 2410 to 3703), are indicated. Adapted from Fauquet et al. (2005), Figure 2 on p. 617.
Search WWH :