Abstract
Evidence has consistently indicated that activation of sphingomyelinases and/or ceramide synthases and the resulting accumulation of ceramide mediate cellular responses to stressors such as lipopolysaccharide, interleukin 1P, tumor necrosis factor a, serum deprivation, irradiation and various antitumor treatments. Recent studies had identified the genes encoding most of the enzymes responsible for the generation of ceramide and ongoing research is aimed at characterizing their individual functions in cellular response to stress. This topic discusses the seminal and more recent discoveries in regards to the pathways responsible for the accumulation of ceramide during stress and the mechanisms by which ceramide affects cell functions. The former group includes the roles of neutral sphingomyelinase 2, serine palmitoyltransferase, ceramide synthases, as well as the secretory and endosomal/lysosomal forms of acid sphingomyelinase. The latter summarizes the mechanisms by which ceramide activate its direct targets, PKC^, PP2A and cathepsin D. The ability of ceramide to affect membrane organization is discussed in the light of its relevance to cell signaling. Emerging evidence to support the previously assumed notion that ceramide acts in a strictly structure-specific manner are also included. These findings are described in the context of several physiological and pathophysiological conditions, namely septic shock, obesity-induced insulin resistance, aging and apoptosis of tumor cells in response to radiation and chemotherapy.
Introduction
Cells and organisms have developed various strategies to deal with adverse changes in their environment. Cellular insult by infectious agents, toxins, nutrient deprivation, or genotoxic stress produces a coordinated systemic response (generally referred to as inflammation), which is aimed at neutralization ofthe insult and initiation oftissue repair. The first line of defense at systemic level is stimulation of the innate immune response, which consists of regulated production of inflammatory mediators, including cytokines like IL-1P, TNFa and IL-6. The typical cellular response to environmental stressors includes the induction of cellular apoptosis (in response to either genotoxic stress or cytokines like TNFa), growth arrest (during nutrient deprivation), increased eicosanoid production, cell migration and adhesion (in the presence of infectious agent). While acute inflammation is protective for the organisms, excessive and long-standing inflammation is harmful and underlies diseases like septic shock, atherosclerosis, asthma, rheumatoid arthritis and inflammatory bowel disease.
Diverse signaling pathways mediate cellular response to stress. The sphingolipid second messengers ceramide, ceramide-1-phosphate, sphingosine and sphingosine-1-phosphate play important role as mediators in many of these pathways. This topic is focused on the role of ceramide in cellular stress response and the mechanisms of its generation and action.
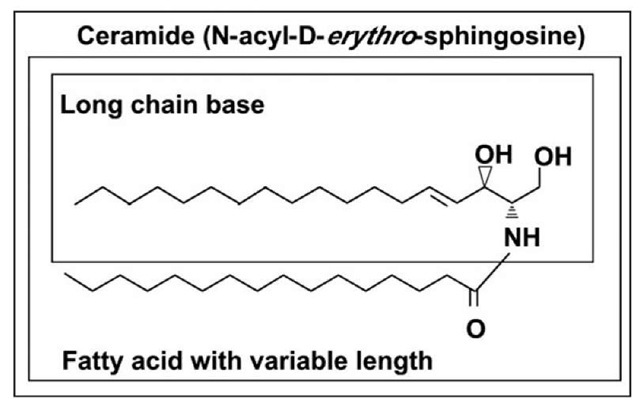
Figure 1. Structure of ceramide.
Chemical Structure and Biophysical Properties of Ceramide
Ceramides (Fig. 1) form the hydrophobic backbone of all complex sphingolipids1 and consist of a long chain sphingoid base and amid linked fatty acid which is either saturated or unsaturated and vary in length from two to 28 carbon atoms. In mammalian cells, the most commonly found ceramides have D-erythro-sphingosine and a saturated fatty acyl chain of 16 carbon atoms and are among the most hydrophobic lipids in the membrane. Free ceramide has a very low critical micellar concentration (cmc < 10~10M) and cannot exist in aqueous solutions.2 Nevertheless, ceramides are still considered as amphiphiles because the hydroxyl group at the first carbon and the amide bond are hydrophilic moieties. Dihydroceramide differs from ceramide inasmuch as the latter contains a trans 4, 5 double bond, which is essential for some of the bioactive roles of ceramide.
The structural features in the ceramide molecule that are required for its biological properties are not well understood, however the two hydroxyl groups, amid group protons and the trans-double bond seems to be involved.3 A network of intramolecular hydrogen bonds involving the OH and NH groups establishes a unique conformation arrangement of ceramide molecule.
Changes in Ceramide Mass during Stress
The basal ceramide concentrations in the cells are low and may change during cellular differentiation or progression through the cell cycle. Various inducers of cellular stress however lead to an accumulation of ceramide that promotes apoptotic, inflammatory and growth inhibitory signals and also mediates the onset of a specific response. The data in Table 1 illustrate what is the magnitude of changes in ceramide levels under various conditions of stress. These reported differences however, are likely to be an underestimation since they are measured in total cell preparations and not in the specific membrane fractions where ceramide was generated. Furthermore, with the exception of mass spectrometry—based assays, the most commonly used methods of ceramide quantification do not separate the individual ceramide species.4 This might be important as the impact ceramide has on cell functions seemingly depends on the type of fatty acid attached to the sphingoid base. Recent data indeed support the notion that the different biological effects caused by ceramide may be mediated by distinct molecular species of the lipid, underscoring the necessity to evaluate changes in ceramide content in a structure-specific manner.
A comparison of the magnitudes of ceramide accumulation observed under various treatments reveals that the changes are more robust in response to chemotherapeutics or irradiation. As the outcome of these treatments has been the induction of cell death via apoptosis, some investigators suggest that ceramide plays the role of a gauge that senses the level of cell injury and depending on downstream factors determines specific biological outcome.5
Table 1. Changes in ceramide content in response to stress and during some pathophysiological conditions
Table 1.
|
|
Control |
Treatment |
Assay |
Tissue/Cell Type |
Ref. |
|
ER stress |
5 pmol/mg Pr |
9 pmol/mg Pr |
Mass Spec |
INS-1 cells |
44 |
|
Ionizing |
100% |
700% of control |
DAGK |
OLG |
45 |
|
radiation |
100% |
320% of control |
Labeling |
CHK cells |
46 |
|
0.95 nmol/106 cells |
1.25 nmol/106 cells |
DAGK |
BAEC |
47 |
|
|
6-12 nmol/ml |
9-18 nmol/ml |
HPLC |
Human serum |
48 |
|
|
100% |
250% of control |
Labeling |
Keratinocytes |
49 |
|
|
10 pmol/106 cells |
50 pmol/106 cells |
Mass Spec |
Jurkat cells |
50 |
|
|
100% |
130% of control |
DAGK |
HEK293 |
51 |
|
|
100% |
140% of control |
DAGK |
Jurkat T-cells |
51 |
|
|
10 pmol/nmol LP |
17 pmol/nmol LP |
Mass Spec |
MCF-7 |
52 |
|
|
100% |
320% of control |
DAGK |
HeLa |
53 |
|
|
100% |
140% of control |
Labeling |
U937 cells |
54 |
|
|
100% |
900% of control |
DAGK |
Human platelets |
54 |
|
|
Diabetes/ |
0.5 pmol/^g Pr |
1.0 pmol/^g Pr |
DAGK |
Rat heart tissue |
55 |
|
obesity |
12 pmol/mg tissue |
23 pmol/mg tissue |
Mass Spec |
Muscle |
56 |
|
25 pmol/mg tissue |
40 pmol/mg tissue |
DAGK |
Muscle |
57 |
|
|
0.2 nmol/mg tissue |
0.3 nmol/mg tissue |
DAGK |
Liver |
57 |
|
|
15 pmol/mg |
23 pmol/mg |
Mass Spec |
Muscle |
58 |
|
|
100% |
No change |
Mass Spec |
Liver |
58 |
|
|
3.5 pmol/^l |
9 pmol/^l |
Mass Spec |
Serum |
58 |
|
|
0.5 nmol/mg Pr |
1.5 nmol/mg Pr |
HPLC |
Liver |
59 |
|
|
2.4 nmol/ml |
3.1 nmol/ml |
Mass Spec |
Human plasma |
60 |
|
|
120 pmol/150^ |
190 pmol/150^ |
Mass Spec |
Mouse Serum |
61 |
|
|
380 pmol/mg Pr |
580 pmol/mg Pr |
Mass Spec |
Epididymal fat |
61 |
|
|
300 pmol/mg Pr |
550 pmol/mg Pr |
Mass Spec |
Subcutaneous fat |
61 |
|
|
650 pmol/ml |
1400 pmol/ml |
Mass Spec |
Mouse plasma |
62 |
|
|
310 pmol/mg Pr |
210 pmol/mg Pr |
Mass Spec |
Epididymal fat |
62 |
|
|
5 nmol/g |
8 nmol/g |
Mass Spec |
Subcutaneous fat |
63 |
|
|
0.6 nmol/mg Pr |
1.2 nmol/mg Pr |
DAGK |
Pancreatic islets |
64 |
|
|
250 pmol/mg Pr |
180 pmol/mg Pr |
Mass Spec |
Retina |
65 |
|
|
Aging |
100% |
170% of control |
Mass Spec |
Cerebral cortex |
66 |
|
100% |
220% of control |
DAGK |
Heart |
67 |
|
|
175 pmol/^g DNA |
250 pmol/^g DNA |
Mass Spec |
Adipose tissue |
68 |
|
|
0.45% of total LP |
0.8% of total LP |
HPLC |
Liver |
69 |
|
|
1.3 nmol/mg Pr |
1.7 nmol/mg Pr |
HPLC |
Rat hepatocytes |
70 |
|
|
0.5 pmol/mg Pr |
1.1 pmol/mg Pr |
DAGK |
Endothelium |
71 |
|
|
3.5 pmol/nmol LP |
6.0 pmol/nmol LP |
DAGK |
WI-38 HDF |
72 |
|
|
Alzheimer’s |
27 nM/ml |
52 nM/ml |
DAGK |
CSF |
73 |
|
disease |
100% |
175% of control |
Mass Spec |
Brain |
74 |
|
2.5 nmol/mg Pr |
9 nmol/mg Pr |
Mass Spec |
Temporal cortex |
75 |
|
|
2.0 nmol/mg Pr |
8 nmol/mg Pr |
Mass Spec |
Cerebellum |
75 |
|
|
Heat stress |
1.5 pmol/nmol LP |
3.8 pmol/nmol LP |
DAGK |
Molt-4 |
76 |
Table 1.
|
|
Control |
Treatment |
Assay |
Tissue/Cell Type |
Ref. |
|
Oxidative |
100% |
800% of control |
DAGK |
Human primary |
77 |
|
stress |
100% |
200% of control |
DAGK |
OLG |
78 |
|
100% |
170% of control |
DAGK |
Cerebral cortex |
79 |
|
|
100% |
200% of control |
DAGK |
PC12 |
80 |
|
|
100% |
350% of control |
Mass Spec |
Airway epithelium |
81 |
|
|
100% |
140% of control |
Labeling |
APRE-19 |
82 |
|
|
100% |
180% of control |
DAGK |
SMC |
83 |
|
|
HAEC |
|||||
|
Car- |
210.5 pmol/mg Pr |
305.9 pmol/mg Pr |
DAGK |
Mouse heart |
84 |
|
diovascular |
100% |
140% of control |
Mass Spec |
Mouse heart |
85 |
|
disease |
135 nmol/ml |
220 nmol/ml |
Mass Spec |
Plasma |
86 |
|
Anti-cancer |
100% |
1000% of control |
Labeling |
LnCaP |
87 |
|
therapies |
100% |
200% of control |
Labeling |
PC-3 |
87 |
|
7 pmol/nmol LP |
70 pmol/nmol LP |
DAGK |
MDA-MB 231 |
88 |
|
|
100% |
220% of control |
Labeling |
MDA-MB 468 |
89 |
|
|
100% |
350% of control |
Labeling |
MCF-7 |
89 |
|
|
1% of total LP |
10% of total LP |
Labeling |
MCF-7 |
90 |
|
|
100% |
900% of control |
Labeling |
MCF-7 |
91 |
|
|
100% |
180% of control |
Labeling |
BT-20 |
92 |
|
|
100% |
300% of control |
Labeling |
MDA-MB 231 |
92 |
|
|
100% |
650% of control |
Labeling |
MDA-MB 468 |
92 |
|
|
100% |
300% of control |
Labeling |
Hs-578T |
92 |
|
|
100% |
380% of control |
Labeling |
T47D |
92 |
|
|
100% |
620% of control |
Labeling |
MCF-7 |
92 |
|
|
100% |
700% of control |
Labeling |
HL-60/VCR |
93 |
|
|
100% |
220% of control |
Labeling |
U-937 |
93 |
|
|
100% |
220% of control |
Labeling |
CHLA-90 |
94 |
|
|
0.6 ng/106 cells |
1.8 ng/106 cells |
DAGK |
JHU-022 |
95 |
In some cases numerical values were taken from graphically represented data and were rounded to the nearest whole number. Abbreviations: General: CSF, cerebrospinal fluid; DAGK, diacylglycerol Kinase; LP, lipid phosphate; M0, primary macrophages; OLG, oligodendrocytes; Pr, Protein. Cells lines: A7r5, rat embryonic thoracic aorta smooth muscle; APRE-19, human retinal pigment epithelium; BAEC, bovine aortic endothelium; BT-20—human breast carcinoma; CG4, oligodendrocyte progenitor; CHK, Chinese hamster kidney; CHLA-90, human neuroblastoma; HAEC, human airway epithelial; HDF, human diploid fibroblasts; HEK 293, human embryonic kidney; HeLa, cervical carcinoma; HL60/VCR, drug resistant human leukemia; Hs-578T, breast carcinoma; HUVEC, human umbilical vein endothelium; INS-1, rat insulinoma; JHU-022, squamous cell carcinoma; Jurkat, T-lymphocytes; L929, mouse fibroblasts; MDA-MB 231, human Caucasian breast adenocarcinoma; MDA-MB 468, human Black breast adenocarcinoma; MCF-7, human breast adenocarcinoma; MIN6, murine beta cells; Molt-4; human acute lymphoblastic leukemia; LnCaP, human prostate carcinoma; NRK-52E, rat kidney epithelium; NT-2, human neuronal precursor; PC3, human prostate carcinoma, PC12, rat adrenal medulla carcinoma; SK2, hybridoma; SP-1, mammary intraductal adenocarcinoma; T47D, human ductal breast epithelial tumor; TNP-1, human acute monocytic leukemia; U87, human astrocytoma; U937, human leukemic monocyte lymphoma.