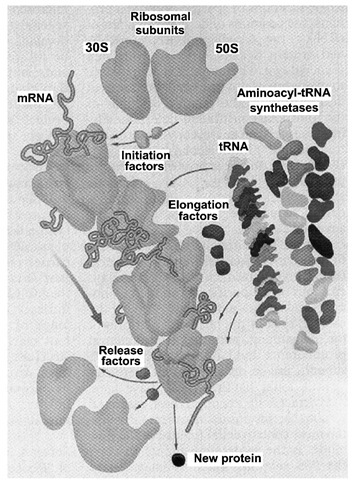
Many cellular components play a role in protein synthesis (Fig. 1). Even in relatively simple bacteria, translation of a single polypeptide from its genetic message requires dozens of participants—proteins, RNAs, and nucleotides—working together as carriers, catalysts, energy sources, and cofactors. Peptide bond formation takes place rapidly at the ribosome, with as many as 40 amino acids per second joined to a growing polypeptide chain. Yet coupled with the need for speed is the requirement for accuracy. The misincorporation of a single amino acid could have drastic effects on the structure or function of a newly synthesized protein. However, protein synthesis is accurate, with errors occurring on the average only once in 10,000 peptide bonds formed.
FIGURE 1 Bacterial protein synthesis. This cartoon summarizes the numerous components necessary for translation of an mRNA into its corresponding polypeptide product. Translation occurs at the ribosome, and is initiated at an AUG start codon by protein initiation factors. Aminoacyl-tRNA synthetases attach amino acids to tRNA adaptor molecules in highly specific enzymatic reactions. Each elongation cycle adds an amino acid to the growing polypeptide chain according to base-pairing interactions between each trinucleotide codon of the mRNA and the anticodon of the matching aminoacylated tRNA. Elongation factors facilitate aminoacyl tRNA selection and, following peptide bond formation, translocation of the peptidyl-tRNA and mRNA. When a stop codon is reached, release factors trigger hydrolysis of the newly synthesized protein and dissociation of ribosomal subunits.
Formation of peptide bonds linking together amino acids could theoretically occur such that random sequences are generated. Some of these sequences could result in a polypeptide that has a useful function. However, transfer of genetic information from one generation to the next requires a systematic and reproducible mechanism for generating defined sequences. Polypeptide formation as we know it today is template-directed, with the messenger RNA (mRNA) copy of a gene providing the text to be deciphered into the protein product.
The simplest linkbetween nucleic acid andprotein components would have been a code with a one-to-one correspondence where each nucleotide dictated a particular amino acid. With only four nucleotides making up the information storage in cells, the resulting proteins synthesized in such a scenario would be limited to those having 4 different amino acids. Even a code of two nucleotides per amino acid would allow for only 16 amino acids. The standard genetic code instead makes use of trinucleotide sequences called codons; these 64 codons are able to determine fully the 20 amino acids used in protein synthesis and also include start and stop codons (Table I).
TABLE I The Standard Genetic Code
|
First position |
Third position |
||||||
|
U |
C |
A |
G |
||||
|
U |
UUU |
Phe |
UCU |
Ser |
UAU Tyr |
UGU Cys |
U |
|
UUC |
Phe |
UCC |
Ser |
UAC Tyr |
UGC Cys |
C |
|
|
UUA |
Leu |
UCA |
Ser |
UAA Stop |
UGA Stop |
A |
|
|
UUG |
Leu |
UCG |
Ser |
UAG Stop |
UGG Trp |
G |
|
|
C |
CUU |
Leu |
CCU |
Pro |
CAU His |
CGU Arg |
U |
|
CUC |
Leu |
CCC |
Pro |
CAC His |
CGC Arg |
C |
|
|
CUA |
Leu |
CCA |
Pro |
CAA Gln |
CGA Arg |
A |
|
|
CUG |
Leu |
CCG |
Pro |
CAG Gln |
CGG Arg |
G |
|
|
A |
AUU |
Ile |
ACU |
Thr |
AAU Asn |
AGU Ser |
U |
|
AUC |
Ile |
ACC |
Thr |
AAC Asn |
AGC Ser |
C |
|
|
AUA |
Ile |
ACA |
Thr |
AAA Lys |
AGA Arg |
A |
|
|
AUG |
Met3 |
ACG |
Thr |
AAG Lys |
AGG Arg |
G |
|
|
GUU |
Val |
GCU |
Ala |
GAU Asp |
GGU Gly |
U |
|
|
GUC |
Val |
GCC |
Ala |
GGC Gly |
C |
||
|
GUA |
Val |
GCA |
Ala |
GAA Glu |
GGA Gly |
A |
|
|
Val |
GCG |
Ala |
GAG Glu |
GGG Gly |
G |
||
a The AUG codon specifies the start of protein synthesis as well as internal methionine residues.