Introduction
The complexities of neurological diseases make it difficult to develop effective methods to treat them. In addition, it is difficult for the drugs used in neurological diseases to reach target cells at a sufficient concentration due to the blood-brain barrier, which may lead to side-effects in other organs, even though newly developed drugs are targeted directly to the origin of the diseases. Nevertheless, as the field of gene delivery gradually matures, new ways to deliver therapies for diseases become possible, and there are reasons to believe that gene therapy would be an optional treatment for neurological diseases.
There are two main classes of vectors used for gene delivery: nonviral vectors and viral vectors (Gardlik et al., 2005). As viral vectors display a higher efficiency and specificity than nonviral vectors, they have been developed as therapeutic gene transfer carriers for various organ systems throughout the body in which the nervous system is involved (Glorioso et al., 2003). The four major viral vector systems used for neurological diseases are recombinant lentivirus (LV), adenovirus (Ad), herpes-simplex virus (HSV), and recombinant adeno-associated virus (rAAV) (Manfredsson et al., 2010). However, the utilization of these vectors is limited by safety concerns because of their pathogenicity (Zhang et al., 2010).
Foamy viruses (FVs), widespread, complex retroviruses with a large genome approximately 12-13 kb, are syncytial viruses containing two different promoters termed IP (internal promoter) and P (promoter) (Löchelt et al., 1995). Due to the specific structure of the viruses and their innocuousness in both human and primate infections, FV vectors have potential as a safe and capable vehicle with a larger capacity for transgenes (Williams, 2008). It has been reported that FV are potentially safer than gammaretroviruses and lentiviral vectors (Trobridge et al., 2009). In this review, we discuss the use of FV vectors as a safe and efficient carrier to deliver therapeutic genes for neurological disorders and other diseases.
Characteristics of foamy virus (Figure 1)
Construction
FVs were first described 50 years ago and are widely distributed in the natural world (Loh et al., 1992). They have been isolated from several mammals, including nonhuman primates, cattle, cats, and horses.
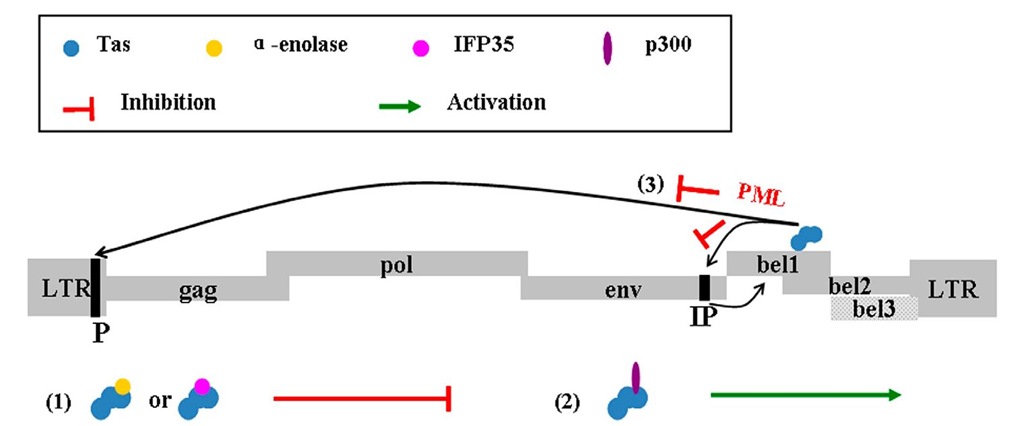
(1) IFP35 can interact with the bovine Tas (BTas) and arrest the replication of BFV; and Bel-1 interacts with a-enolase, then represses the activation of LTR and IP 1 (2) Tas can be acetylated by p300, and activate the virus replication 1 (3) PML inhibits the binding of Tas to IP and LTR.
Fig. 1. The genomic organization of FVs and Tas of FVs interacted with other proteins in cells
Once infected by FVs, life-long latency of the infected cell is obtained (Maxine et al., 2000). FVs contain the largest genome and possess the most complex gene structure of all the retroviruses. For example, the human foamy virus (HFV, also called primate foamy virus (PFV)) contains LTR (long terminal repeat) sequences at the 5′and 3′ ends of its genome, and there are three structural genes between these two LTRs that are found in all viral genomes: gag, pol, and env (Saïb et al., 1995). Several ORFs (open reading frames), often two or three, are located between the env and the LTR at the 3′end of all sequenced FVs, which encode autologous transactivators, including Tas, an important transactivator (Linial, 2000). The Tas protein is required by both the promoter (P) in the U3 region of the 5′ LTR and another unique internal promoter (IP) at the end of the env gene for high-level expression (Linial, 2000).
Promoters
LTR
Long terminal repeats (LTRs), located at the 5′ and 3′ end of the viral genome, are noncoding regions that exist in all types of retroviruses. The LTR is required for the integration of the viral genome into the host genome and is related to the replication, kinetics, tissue tropism, and pathogenicity of the virus. A U3-R-U5 (unique 3′ subregion-repeat subregion-unique 5′ subregion) structure forms the LTR. The LTR promoter, containing a component of the U3-R-U5 region, is a traditional retroviral promoter. The prototypical HFV U3 region is approximately 777 nt and includes a TATA box, transcription regulatory signals and several BREs (Bel1 transactivator response elements) that are located at ~89-116 nt, ~284-327 nt, ~348-360 nt, ~418-454 nt and ~506-559 nt, respectively (Lee et al., 1993; Verdin et al., 1995). As there are a number of BRE regions, Choy, Kerr et al. suggested that the Bel1 regions regulate gene expression by integrating with intracellular transcription factors and transcription initiation complexes without binding directly to BREs and act as enhancers that work together to make a change (Lee et al., 1993; Choy et al., 1993; Kerr et al., 1993; Yang et al., 1997; Löchelt et al., 1994)). The negative control region in the RU5 inhibits the gene expression of HFV, and the viral mRNA R-U5 forms a stable secondary structure (Nabel et al., 1987). There is a splice donor site (SD) 51 nt downstream of the transcription site of the R region, where the posttranscriptional splicing of the gag, pol, env, and bel genes is initiated by the 5′ LTR promoter start site (Li et al., 1999).
IP
All FVs possess a promoter in the U3 region of the LTR. In addition to the LTR promoter P, they also contain another unique internal promoter, termed IP (Löchelt et al., 1993). The internal promoter is located at the end of env gene. These two promoters require Tas for high-level expression (Löchelt et al., 1994). Compared with the LTR promoter, Tas has a higher affinity for the IP (Kang et al., 1998). In the early stage of viral infection, the IP initially regulates the expression of Tas transcription through an LTR cis-acting effect and the participation of as yet unknown cytokines. Next, Tas affects the IP through its self-activating mechanism to elevate its synthesis level; when the Tas protein reaches a certain level, the activated LTR initiates the transcription and expression of the structural gene and performs transformation of gene transcription and expression from the early non-structural gene to the late structural gene.
Structural genes
Following the 5′ LTR of FVs, there are three structural genes (gag, pol, env). The newly synthesized 74/78 kDa precursor protein encoded by the gag gene is located in the nuclei of the infected cells (Schliephake et al., 1994), and it is cleaved into the matrix, capsid, and nucleocapsid structural protein by the viral protease (Saïb et al., 1995). The nucleocapsid (NC) domain of Gag protein lacks Cys-His boxes found in all other retroviral Gag proteins, but its C terminus possesses three glycine-arginine motifs (GR boxes) that are required for RNA packaging (Stenbak et al., 2004; Yu et al., 1996; Maurer et al., 1998). The pol gene overlaps the gag gene, and its product does not contain any Gag protein determinant; however, unlike other conventional retroviruses, there is no Gag-Pol fusion protein synthesized in the course of viral transcription, but the underlying mechanism remains unknown (Stenbak et al., 2004; Yu et al., 1996). The precursor protein encoded by the env gene, which is approximately 130 kDa, is cleaved into the SU and TM subunits during the process of maturation (Saïb et al., 1995; Aguzzi, 1993). Linial et al. found that the extracellular virions of HFV require the Env protein for further production but that the formation of intracellular particles seems to be independent of the Env protein (Baldwin et al., 1993). Unfortunately, the role of Gag, Pol, and Env in the assembly of the FVs remains uncertain.
Regulatory genes
There are several ORFs encoding accessory genes or regulatory genes that are located between the env and the LTR and are therefore named bel genes. These ORFs encode autologous transactivators, including bel-1, bel-2, bel-3 and bet.
Tas
The regulatory gene bel-1, which is approximately 902 bp in length, is located in the nucleus of expressing cells and is essential for HFV replication. FVs transactivators of different genera have been named differently; the simian FVs transactivator, for example, was termed taf. However the First International Foamy Virus Meeting determined that these transactivators, including Taf and Bel1, should be designated Tas (Zou et al., 1996). The activity of the functional IP and P of FVs depends on this transactivator. Transcription starts from the integration of Tas and the IP, leading to transactivation of the IP. However, the LTR promoter will not be activated until the level of Tas reaches a certain level (Linial, 2000). In view of the fact that the Tas protein plays an important role in the transcription and replication of FVs, it has received increasing attention and interest among virologists.
The interactions between Tas and other cellular molecules are complex. For example, Tas can be acetylated by p300 and PCAF (Bodem et al., 2007). Recently, an interaction between Tas and transcriptional coactivators, such as HATs, was reported. Studies show that only acetylated Tas enhances viral transcription (Bodem et al., 2007). Interferon-induced proteins (IFPs) complete multiple functions through various interferon signals. IFP35, which is approximately 35 kDa, is a member of the IFP family. It interacts with the bovine foamy virus Tas regulatory protein, and its overexpression disturbs viral gene transcription and replication of the bovine foamy virus Tas protein. Furthermore, IFP35 plays a crucial role in the interferon-induced anti-foamy virus host response (Tan et al., 2008). First discovered by Lohman and Mayerhof in 1934 (Lohman et al., 1934), α-enolase was later shown to be a highly conserved enzyme that catalyses the transformation of 2-phosphoglyceric acid to enolphosphopyruvate in the Embden-Meyerhof-Parnas pathway (Wold, 1971). α-Enolase interrupts the transactivation of Tas with the promoters during viral transcription (Wu et al., 2009). The promyelocytic leukemia protein (PML) is a cell growth and tumor suppressor gene, which is expressed in the nucleus and whose expression is increased significantly in the response to interferon (IFN) (Wang et al., 1998). Regad et al. found that PML inhibits the binding of Tas to the IP and LTR by direct interaction with Tas (Regad et al., 2001).
Bet
Bet is a second IP regulating protein. Generated by the combination of the spliced mRNA of the Tas region and the Bel2 ORF (Linial, 2000), overexpression of Bet prevents the virus from infecting the host and inhibits viral integration at the initial stage of viral infection (Bock et al., 1998). A variant RNA can be generated by the LTR promoter, which has the ability to generate Bet protein fragments that, after reverse transcription, could induce the expression of a ATas provirus (Saïb et al., 1993). Because of the deficiency of Tas in the provirus, ATas plays a significant role in the preservation and latency of FVs, which clarifies that the LTR does not initiate the synthesis of viral protein, whereas the expression of Bet is induced by IP (Linial, 2000). Transcription may be different in diverse cell types. Additionally, the combination site of Tas for the internal promoter IP is different from that for the LTR promoter, and the required cytokines are different. High expression of the virus would be suppressed under conditions in which the IP promoter was active in various cell types, yet cytokines, which the LTR promoter requires, exist in a select few cells. The transcription of integral provirus is mainly initiated by IP and will express Tas and Bet but virus will not.
Replication and transcription
As mentioned previously, FVs are a member of the spumavirus genus of the retroviridae family. It is believed that the replication and transcription mechanisms of various family members are similar (Coffin, 1990). However, Yu et al. found that although HFV possesses a replication pathway with features of both retroviruses and hepadnaviruses, there are also unique features contained in the assembly and reverse transcription pathway (Yu et al., 1996).
At an early stage of viral infection, Tas is expressed due to the high level of transcription of IP, which in turn transactivates the expression of IP and LTR, leading to the expression of accessory and structural genes, respectively, inducing late stage viral infection (Löchelt et al., 1994). The Gag-Pol fusion protein is synthesized during the transcription of other retroviruses. However, as discussed above, the pol gene product in foamy viruses does not contain any Gag protein determinant, which means there is no Gag-Pol fusion protein synthesized in the course of viral transcription. Therefore, unknown mechanisms exist in the synthesis of pol gene products. As there are two promoters present in the genome of foamy virus, it is presumed that IP functionally replaces the post-transcriptional regulatory protein of other retroviruses and thus provides a temporal pattern with stage-specific promoters (Mergia, 1994).
Although FVs infection can result in a unique foam-like cytopathic effect and syncytia in cell cultures, they are nonpathogenic in naturally or experimentally infected animals, and the pathogenicity to their hosts remains unknown. However, more and more research groups are attracted to FVs due to the lack of pathogenicity; thus, increased attention has been given to the development of FVs as vectors of target gene delivery.
The development of human foamy virus (HFV) (Figure 2)
Replication-competent HFV
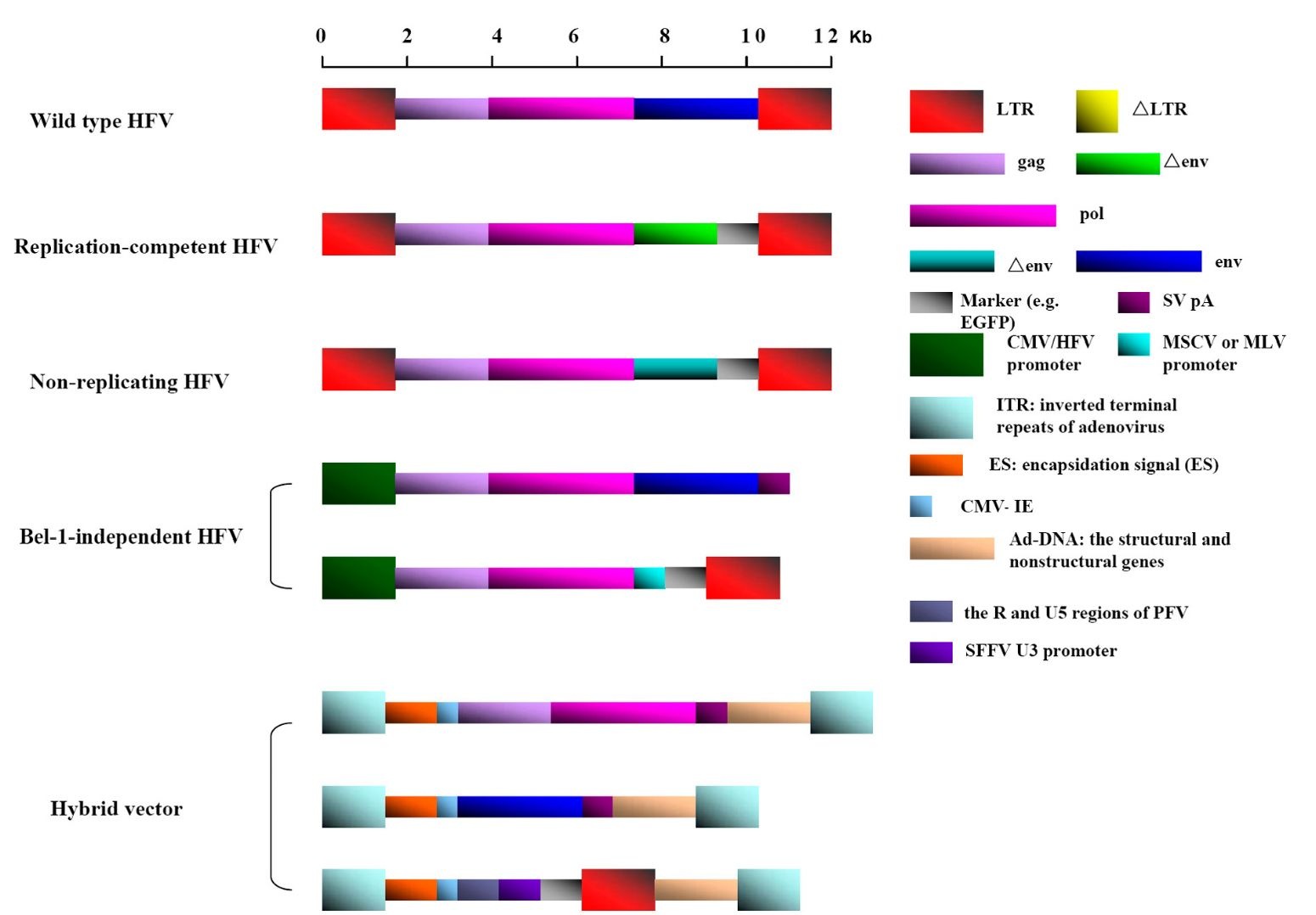
The first described FVs vectors (pFOV-1, pFOV-3 and pFOV-7) were replication-competent. Noncoding sequences in bel-2 and bel-3 were deleted, and two termination codons and a bel-1 splice donor site mutation were introduced into these vectors (Schmidt et al., 1995). The vector pFOV-7, containing the herpes virus type 1 (HSV-1) thymidine kinase (tk), the E.coli cytosine deaminase (cd) and the polynucleotide phosphorylase (pnp) genes, was introduced into tumor cells and promoted their death (Nestler et al., 1997).
Fig. 2. The development of HFV vectors. The replication-competent FV vectors were deleted noncoding sequences in bel-2 and bel-3. The non-replicating FV vectors were constructed by the portion of the env gene replaced by marker gene. The bel-1-independent FV vector system contains relevant packaging plasmid and the vector plasmid carrying exogenous gene. The adenovirus and human foamy virus hybrid vector system express separately the HFV structural genes gag and pol, the HFV structural gene env, and the transgene.