microCosm, TargetScan, and PicTar
The miRBase database links miRNAs to targets using microCosm (http://www.ebi.ac.uk/enright-srv/microcosm/), TargetScan (Lewis et al., 2005; Grimson et al., 2007; Friedman et al., 2009) and PicTar (Lewis et al., 2005; Grimson et al., 2007; Friedman et al., 2009) prediction systems. These are therefore the most currently used, and are detailed below.
microCosm, formerly known as miRBase Targets, predicts miRNA targets in the UTR regions of animal genomes from Ensembl database (Hubbard et al., 2007; Flicek et al., 2008). It uses the miRanda algorithm to calculate a score across the miRNA vs UTR alignment (Enright et al, 2003; John et al, 2004; Betel et al, 2008); the energy for the thermodynamic stability of a miRNA : mRNA duplex is calculated by the Vienna RNA folding routines (http://www.tbi.univie.ac.at/RNA/), and the P-values are computed for all targets following the statistical model implemented in RNAhybrid (Rehmsmeier et al., 2004). The Miranda algorithm (Enright et al., 2003; John et al., 2004) is basically divided into three steps. In the first step the miRNAs are aligned against the 3′ UTR sequences of the targets, allowing for G : U pairs and short indels. The method does not rely on seed matches, but increases the scaling score for complementarity at the 5′ end of the miRNA. The second step computes the ther-modynamic stability of the miRNA : mRNA duplex, and the final step reduces the false-positive rate by considering only targets with multiple sites.
TargetScan was the first algorithm that used the concept of seed matches in target prediction (Lewis et al., 2003, 2005). The method only uses miRNAs conserved across different species to scan corresponding 3′ UTR sequences.
Table 3 Algorithms Developed for Predicting miRNA Targets
|
Algorithm |
Strategy |
Species group |
Authors/year |
|
TargetScan |
RB |
Vertebrates |
Lewis et al., 2003 |
|
TargetScanS |
RB |
Vertebrates |
Lewis et al., 2005 |
|
miRanda |
RB |
Insects (flies), Human |
Enright et al., 2003; John et al., 2004 |
|
Diana-microT |
RB |
Nematodes |
Kiriakidou et al., 2004 |
|
RNAhybrid |
RB |
Insects (flies) |
Rehmsmeier et al. , 2004 |
|
MovingTargets |
RB |
Insects (flies) |
Burgler and MacDonald, 2005 |
|
MicroInspector |
RB |
Any species |
Rusinov et al., 2005 |
|
Nucleus |
RB |
Insects (flies) |
Rajewsky and Socci, 2004 |
|
EIMMo |
RB |
Nematodes, Insects (flies), Vertebrates |
Gaidatzis et al., 2007 |
|
TargetBoost |
BT |
Nematodes, Insects (flies) |
Saetrom et al., 2005 |
|
PicTar |
HMM |
Nematodes, Insects (flies), Vertebrates |
Krek et al., 2005 |
|
RNA22 |
MC |
Nematodes, Insects (flies), Vertebrates |
Miranda et al., 2006 |
|
MicroTar |
PD |
Any species |
Thadani and Tammi, 2006 |
|
PITA |
PD |
Nematodes, Insects (flies), Vertebrates |
Kertesz et al., 2007 |
|
NBmiRTar |
NB |
Metazoa |
Yousef et al., 2007 |
|
miTarget |
SVM |
Metazoa |
Kim et al., 2006 |
|
MiRTif |
SVM |
Metazoa |
Yang et al., 2008 |
|
mirWIP |
E |
Nematodes |
Hammell et al., 2008 |
|
Sylamer |
E |
Metazoa |
van Dongen et al., 2008 |
|
GenMiR++ |
BL, E |
Metazoa |
Huang et al., 2007 |
|
SVMicrO |
SVM, E |
Mammals |
H. Liu et al., 2010 |
|
TargetMiner |
SVM, E |
Human |
Bandyopadhyay and Mitra, 2009 |
|
MirTarget2 |
SVM, E |
Metazoa |
Wang and El Naqa, 2008 |
|
TargetSpy |
BT, E |
Insects (flies), Human |
Sturm et al., 2010 |
The algorithm defines the seed matches as short segments of seven nucleotides that must have a stringent complementarity to the two to eight nucleotides of the mature miRNA. Then, the remaining miRNA sequence is aligned to the target site, allowing for G : U pairs; the free energy to form a secondary structure in the duplex is predicted by a folding algorithm. A Z-score is calculated on the basis of the number of matches predicted in the same target sequence and respective free energies. Finally, the Z-score is used to rank the candidate targets for each species, and each species is processed in the same way.
PicTar uses a machine learning algorithm to rank target sequences using a HMM maximum likelihood score based on three main steps: (1) the seed matches must expand 7 nucleotides starting at position 1 or 2 in the 5′ end of the miRNA; (2) the minimum free energy of miRNA : mRNA duplexes is used to filter the target sites; and (3) the target sites must locate in overlapping positions across the aligned corresponding 3′ UTR sequences. The target sites that pass the three-step filter are then ranked by the HMM model, which calculates the score considering all segmentations of the target sequence into target sites and background, thus allowing the algorithm to account for multiple binding sites for a single miRNA, as well as several miRNAs targeting the same mRNA.
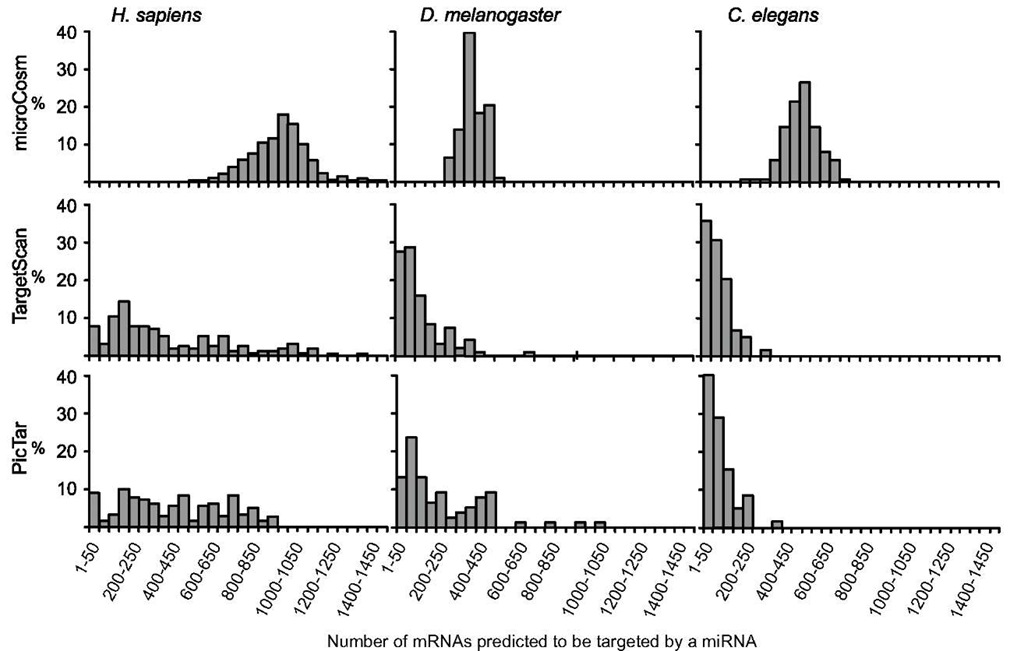
The current target predictions available in the miRBase by microCosm, TargetScan, and PicTar have some degree of overlap and also of discrepancy that can be due to alignment artifacts, different mRNA UTR and miRNA sequences, and intrinsic differences in the algorithms. In an attempt to provide more updated figures for the distribution of gene targets per miRNA and miRNA per gene target, we analyzed the data from target predictions available in the miRBase (Release 16; Sept 2010), comparing D. melanogaster with Homo sapiens and C. elegans. Results show that the three methods give different average numbers of miRNA-binding sites per mRNA target (19.6, 5.8, and 5.0 for MicroCosm, TargetScan, and Pic-Tar, respectively; Figure 7), as well as different numbers of mRNAs targeted by each miRNA (951, 395, and 426 for microCosm, TargetScan, and PicTar, respectively; Figure 8). The distribution of the number of miRNA-binding sites per mRNA target (Figure 7) is relatively similar among the three methods and the three species studied. Conversely, data on the number of mRNA targeted by an miRNA showed remarkable differences depending on the method, regarding not only the average values, but also and especially their pattern of distribution (Figure 8).
miRNA Functions
Insect model species can be studied through powerful genetic and genomic approaches, the paradigm being the fly D. melanogaster. Indeed, the first description of miRNA functions in insects was carried out in this species (Brennecke et al., 2003), by looking at gain-of-function mutants (Lai, 2002; Lai et al., 2005). miRNA functions are currently being demonstrated by mutating the genes coding for the miRNAs under study, overexpressing the miRNA of interest, or silencing it using specific anti-miRNAs, and then studying the resulting phenotype.
Figure 7 Frequency of the number of miRNA-binding sites in the 3′ UTR of target mRNAs in Homo sapiens, Drosophila melanogaster, and Caenorhabditis elegans, calculated with the three prediction methods available in miRBase: microCosm, TargetScan, and PicTar.
Figure 8 Frequency of the number of mRNAs predicted to be targeted a miRNA in Homo sapiens, Drosophila melanogaster, and Caenorhabditis elegans, calculated with the three prediction methods available in miRBase: microCosm, TargetScan, and PicTar.
Predicted targets may also be validated by the above methods, including the quantification of the expression of the given target, as well as using in vitro systems with luciferase reporter target constructs, where binding of the miRNA to the target sequence is detected by luciferase activity and quantified with colorimetry.
In most cases, functions may be suggested by high-throughput sequencing comparisons in different developing stages, in different organs of the same stage, or in different physiological situations. Studies of this type have been carried out in the silkworm B. mori (differences in tissue expression and in different developing stages) (Cao et al., 2008; S. Liu et al., 2010), the pea aphid A. pisum (differences in different morphs) (Legeai et al., 2010), the honey bee A. mellifera (differences between queens and workers) (Weaver et al., 2007), the migratory locust L. migratoria (differences between migratory and solitary phases) (Wei et al., 2009), and the German cockroach B. germanica (differences between metamorphic and non-metamorphic instars) (Cristino et al., 2011). Microarray analysis or detailed studies on the developmental expression profiles of particular miRNAs can also suggest their respective functions (Aravin and Tuschl, 2005; Weaver et al., 2007; He et al., 2008; Yu et al., 2008).
Silencing Dicer-1 expression by RNAi is also a useful approach to studying the influence of the whole set of miRNAs in a given process. This has been achieved in D. melanogaster, either in vivo, showing, for example, that Dicer-1 plays a general role in ovarian development (Jin and Xie, 2007), or in Drosophila cultured cells, where the depletion of Dicer-1 affected the development in both somatic and germ lineages (Lee et al., 2004b). More recently, Dicer-1 depletion by RNAi has been used in the German cockroach, B. germanica, to demonstrate the key role of miRNAs in hemimetabolan metamorphosis (see below).
Regarding the functions of particular miRNAs, the data available indicate that most of them appear to be involved in the fine-tuning of biological processes by modulating a precise dosage of regulatory proteins. Probably, they provide robustness to the whole program of gene expression (Hornstein and Shomron, 2006) and resilience to environmental fluctuations, as in the case of miR-7 studied by Li and colleagues (X. Li et al., 2009). However, as revealed by recent general reviews (Bushati and Cohen, 2007; Jaubert et al., 2007), information is still fragmentary, heavily concentrated in the D. melanogaster model, and focused on a few biological processes, as detailed in the text below and in Table 4, which summarizes cases where the miRNA function has been demonstrated experimentally.
Table 4 Functions of miRNA Demonstrated Experimentally*
|
Function/process |
miRNA |
Target involved |
Authors/year |
|
Cell division of the germinal stem cells |
bantam |
Hatfield et al., 2005; Shcherbata et al., 2007 |
|
|
Cell division of the germinal stem cells |
miR-7, miR-278, miR309 |
Dacapo |
Yu et al., 2009 |
|
Germ-line differentiation |
miR-7 |
bam |
Pek et al., 2009 |
|
Stem cells differentiation |
miR-184 |
Saxophone |
Iovino et al., 2009 |
|
Axis formation in the egg chamber |
miR-184 |
Gurken |
Iovino et al., 2009 |
|
Formation of the head and posterior abdominal segments in the embryo |
miRs-2/13 |
Boutla et al., 2003 |
|
|
Embryo segmentation |
miR-31, miR-9 |
Leaman et al., 2005 |
|
|
Embryo growth |
miR-6 |
Leaman et al., 2005 |
|
|
Formation of embryonic cuticle |
miR-9 |
Leaman et al., 2005 |
|
|
Photoreceptor differentiation |
miR-7 |
Yan |
Li and Carthew, 2005 |
|
Formation of sensory organs |
miR-9a |
Senseless |
Li et al., 2006 |
|
Location of CO2 neurons |
miR-279 |
Nerfin-1 |
Cayirlioglu et al., 2008 |
|
Protection of sense organs from apoptosis |
miR-263a/b |
Hid |
Hilgers et al., 2010 |
|
Muscle differentiation |
miR-1 |
Delta |
Kwon et al., 2005 |
|
Muscle differentiation |
miR-133 |
nPTB |
Boutz et al., 2007 |
|
Growth |
bantam |
Hipfner et al., 2002; Edgar, 2006; Thompson and Cohen, 2006 |
|
|
Tissue growth via insulin receptor signaling |
miR-278 |
Teleman et al., 2006 |
|
|
Growth via insulin receptor signaling |
miR-8 |
U-shaped |
Hyun et al., 2009 |
|
Modulation of ecdysteroid pulses |
miR-14 |
EcR |
Varghese and Cohen, 2007 |
|
Neuromusculature remodeling during metamor- |
let-7 (and miR-100, |
Sokol et al., 2008 |
|
|
phosis |
miR-125) |
||
|
Maturation of neuromuscular junctions during metamorphosis |
let-7 (and miR-125) |
abrupt |
Caygill and Johnston, 2008 |
|
Wing formation |
miR-9a |
dLOM |
Biryukova et al., 2009 |
|
Wing formation |
iab-4 |
Ultrabithorax |
Ronshaugen et al. , 2005 |
|
Regulation of circadian rhythms |
bantam |
clock |
Kadener et al., 2009a |
|
Regulation of brain atrophin |
miR-8 |
atrophin |
Karres et al., 2007 |
|
Anti-apoptotic |
Bantam, miR-2 |
hid |
Brennecke et al., 2003, 2005; Stark et al., 2005 |
|
Anti-apoptotic in D. melanogaster |
miR-14 |
Drice |
Xu et al., 2003 |
|
Anti-apoptotic in Lepidopteran Sf9 cells |
miR-14 |
Kumarswamy and Chandna, 2010 |
|
|
Anti-apoptotic in the embryo |
miR-2, miR-13, miR-11 |
hid, grim, reaper, sickle |
Leaman et al., 2005 |
*All results refer to Drosophila melanogaster, except in the anti-apoptotic action of miR-14, which has been demonstrated also in Sf9 cells of the Lepidopteran Spodoptera frugiperda.
Germ-Line and Stem Cell Differentiation, Oogenesis
In D. melanogaster, cell division of the germinal stem cells (GSC) is under the control of different miRNAs. One of them, bantam, regulates the expression of specific mRNAs in the ovary, being involved in the maintenance of germinal stem cells (Hatfield et al., 2005; Shcherbata et al., 2007). Other miRNAs, like miR-7, miR-278 and miR-309, directly repress Dacapo mRNA through its 3′ UTR, as demonstrated by Yu and colleagues (2009) using luciferase assays. These authors also suggest that bantam and miR-8 regulate Dacapo indirectly, controlling GSC division; moreover, GSC deficient for miR-278 show a mild, but significant, reduction of cell proliferation. Depletion of miR-7 levels in GSC results in a perturbation of the frequency of Cyclin E-positive GSC, although the kinetics of cell division in miR-7 mutant GSC does not become reduced (Yu et al., 2009).
Another miRNA that plays important roles in stem cell differentiation is miR-184. Depletion of miR-184 in D. melanogaster determines that females lay abnormal eggs and become infertile. Stem cell differentiation is impaired due to the increase of Saxophone protein levels. Later, during oogenesis, the absence of mir-184 impairs the axis formation of the egg chamber as a result of altering the expression of Gurken mRNA. In addition, the absence of miR-184 also affects the expression of pair-rule genes required for normal anteroposterior patterning and cellularization of the embryo (Iovino et al, 2009).
Finally, and also in D. melanogaster, miR-7 is involved in germ-line differentiation via maelstrom and Bag-of-marbles (Bam) gene products. Maelstrom regulates Bam via repression of miR-7, by binding to the miR-7 promoter region (Pek et al., 2009); therefore, D. melanogaster mutants for maelstrom overexpress Bam, which leads to a deficient germ-line differentiation. As expected, a reduction in miR-7 expression rescues this phenotype (Pek et al., 2009)