Summary
MicroRNAs (miRNAs) are endogenous, ca. 22-nucleo-tide, single-strand, non-coding RNAs that regulate gene expression by acting post-transcriptionally through base-pairing between the so called "seed" sequence of the miRNA (nucleotides 2-8 at its 5′ end) and its complementary seed match sequence present in the 3′ untranslated region of the target mRNA. Since the discovery of the first miRNAs in the 1990s, a remarkable diversity of miRNAs has been reported in various organisms, including insects, plants, viruses, and vertebrates. Moreover, computational methods have been developed to find new miRNAs as well as mRNA targets. In insects, most miRNAs are involved in modulating a precise dosage of regulatory proteins, thus fine-tuning biological processes like cell proliferation, apoptosis and growth, oogenesis and embryogenesis, nervous system and muscle differentiation, metamorphosis and other morphogenetic processes, and response to biological stress. The miRNA field is still developing, and many questions remain to be solved. Technologies to determine new miRNAs and miRNA targets still need refinement. Further studies are also needed to elucidate the mechanisms regulating miRNA expression, to validate the miRNA targets in vivo, and to establish the complex networks that connect miRNAs, mRNAs, and proteins, and that govern the development and function of cells and tissues.
Introduction: The Big World of Small RNAs
Step by step, some of the old paradigms of molecular biology have been falling away. The most significant of these is the central dogma that "one gene equals one protein." It still holds true that most information flows from DNA to proteins through intermediate RNA molecules, but today it is well known that the transcriptome is much more complex and diverse than the genome, thanks to the interplay of a variety of mechanisms. The most thoroughly studied is alternative splicing; that is, the formation of diverse mRNAs through differential splicing of the same RNA precursor, which gives rise to proteins with distinct features. Another factor accounting for transcriptome diversity in quantitative terms and in time and space is the occurrence of transcription factors, sequence-specific
DNA-binding factors that usually bind to the promoter region of target genes, thereby activating or repressing their transcription. However, to understand thoroughly the dynamics of the proteome, we have to account for the unknown mechanisms other than simply protein-coding genes and transcription factors. At least, non-coding RNAs (ncRNAs) must also be taken into account in order to have a more complete picture of what is really happening in genomic regulation.
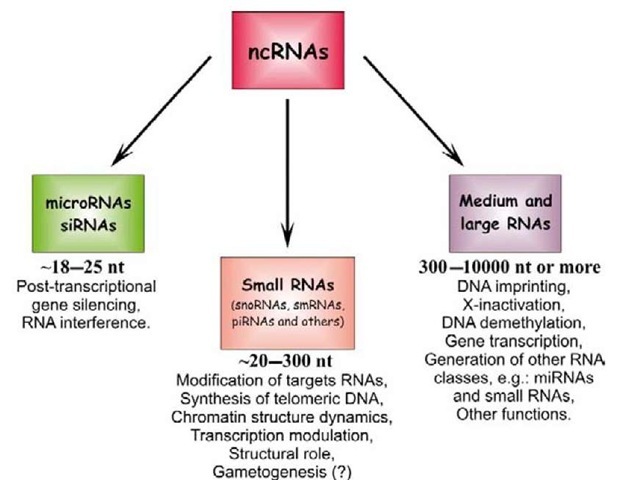
ncRNAs form a heterogeneous group of RNA molecules that are classified into three categories according to their length and function. They range in length from 18 to 25 nucleotides for the group of very small RNAs, which includes short interfering RNAs (siRNAs) and microRNAs (miRNAs); from 20 to 200 nucleotides for the group of small RNAs, which usually play the role of transcriptional and translational regulators; and, for the group of medium and large RNAs, up to (and even beyond) 10,000 nucleo-tides, which are involved in other processes, as detailed in Figure 1 (Costa, 2007). This topic deals with the very small RNAs, and, more specifically, with miRNAs.
Figure 1 Types of non-coding RNAs (ncRNAs) classified according to their length and functions: very small RNAs – microRNAs and small interfering RNAs (siRNAs); small RNAs; and medium and large RNAs. The corresponding established functions for each type are also indicated. snoRNAs, small nucleolar RNAs; smRNAs, small modulatory RNAs; piRNAs, Piwi-interacting RNAs.
The history of siRNAs and miRNAs began in the late 1980s, when Jorgensen and colleagues were studying the role of chalcone synthase in the biosynthetic pathway of anthocianin in plants. Anthocianin gives a violet color to petunias, and Jorgensen’s team overexpressed chal-cone synthase in search of petunias with a deeper violet color. However, they unexpectedly obtained whitish flowers because the expression of chalcone synthase in these transgenic whitish petunias was some 50 times lower than in the wild type, thus suggesting that transgenic chalcone synthase had suppressed the endogenous gene (Jorgensen, 1990). Three years later, but in the field of developmental biology and working on the nematode Caenorhabditis elegans, Lee and colleagues (1993) discovered two lin-4 transcripts, where the smaller, with ca. 21 nucleotides, was complementary to seven repeated sequences in the 3′ UTR of the mRNA of the heterochronic gene lin-14, which had been identified two years earlier.
These two disparate studies converged in 1998, when Fire and colleagues (1998), also working in C. elegans, discovered that the administration of a double-stranded RNA (dsRNA) with a strand complementary to a fragment of an endogenous mRNA can block this mRNA. This phenomenon is now known as RNA interference (RNAi), and its action is mediated by siRNAs of ca. 22 nucleotides that derive from dsRNA (Belles, 2010). A year later, while studying post-transcriptional gene silencing as a mechanism of antiviral defense, Hamilton and Baulcombe (1999) noticed the occurrence of antisense viral RNA of ca. 25 nucleotides in virus-infected plants. Hamilton and Baulcombe observed that these small RNAs were long enough to convey sequence specificity, and pointed out that they might be key determinants of the gene silencing phenomenon. Further contributions showed that dsRNA-induced mRNA degradation was always mediated by RNAs of 21-23 nucleotides, thus leading researchers to investigate the endogenous source of these small RNAs. Finally, in 2001, three groups working independently (Lagos-Quintana et al., 2001; Lau et al., 2001; Lee and Ambros, 2001) described miRNAs as a novel family of small (ca. 22 nucleotides) endogenous RNAs that is diverse in sequence and temporal expression, evolutionarily widespread, and involved in regulating gene expression.
RNAi and siRNAs
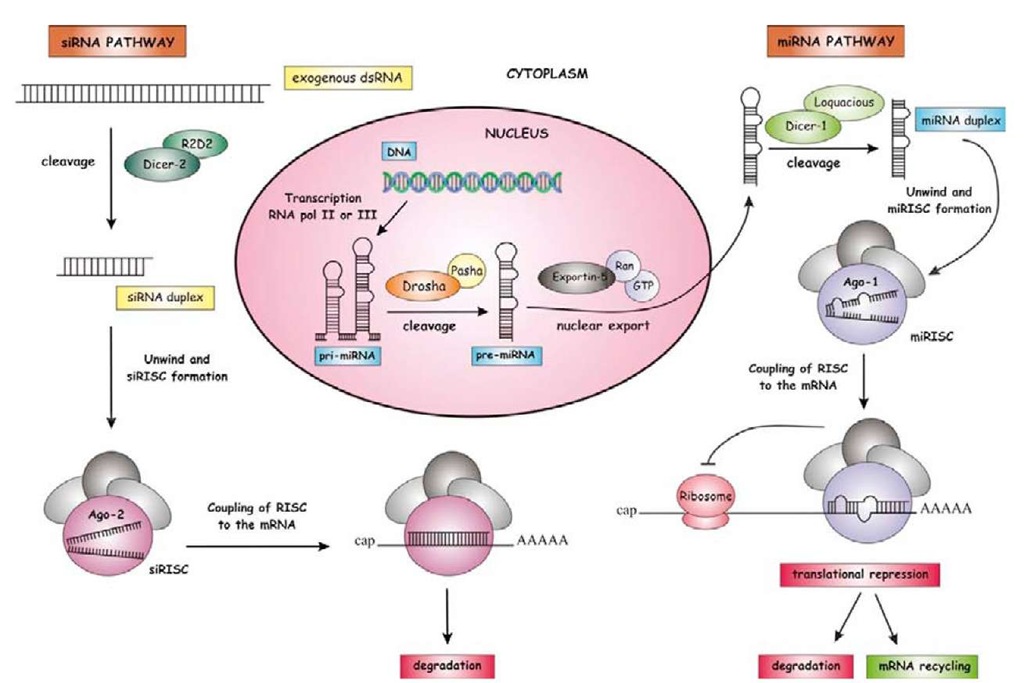
The discovery of RNAi in C. elegans (Fire et al., 1998) was later extended to other animal groups, namely insects, and the basic mechanisms involved in their action on mRNAs were unveiled step by step in a few years. The biochemical machinery and the effects of RNAi in insects have been the subject of a recent review (Belles, 2010), and we will not deal with them in detail here. In essence, when a long and exogenous dsRNA is delivered to the insect, it is cleaved by the enzyme Dicer-2 into siRNA duplexes of ca. 22 nucleotides. These siRNAs then unwind, and single-strand siRNAs bind to Argonaute-2 protein (Ago-2) and assemble into the so-called RNA-induced silencing complex (RISC). The RISC, guided by the siRNA, couples to the target mRNA and degrades it (Belles, 2010). Indeed, the mechanisms of generation and the action of siRNAs are very similar to those of miRNAs (Figure 2), with the latter detailed in the following sections.
The advent of RNAi represented a new paradigm in insect functional genomics, because it opened the door for studying non-drosophilid species – that is, species that cannot be genetically transformed, at least not very easily. RNAi experiments are relatively simple, consisting of conveying a dsRNA with a strand complementary to a fragment of the target mRNA to the animal or to cells incubated in vitro. After assessing that target mRNA levels have lowered, the study of the phenotype unveils the functions associated with the target mRNA. Experiments can be carried out in vitro, where the easiest system involves the incubation of cells with dsRNA added to the medium and then studying the cell behavior; and in vivo, where the most straightforward approach consists in delivering the dsRNA to the chosen insect stage (from egg to adult) of the experimental specimen and then examining the resulting phenotype. The approaches in vivo have afforded the most spectacular results on insect functional genomics (Belles, 2010).
Kennerdell and Carthew (1998) were the first to use RNAi in vivo in insects, studying the genes frizzled and frizzled 2 in the fly Drosophila melanogaster. A year later, Brown and colleagues (1999) carried out functional studies of Hox genes in the flour beetle Tribolium castaneum, and the following year used RNAi for the first time in a hemi-metabolan species, the milkweed bug Oncopeltus fasciatus, on which Hughes and Kaufman (2000) studied Hox gene functions as well. In 2006, RNAi in vivo was used for the first time on a phyllogenetically very basal insect, the German cockroach Blattella germanica (Ciudad et al., 2006; Martin et al. , 2006), which has been shown to be one of the species most sensitive to RNAi. In the past few years an explosion of papers has truly changed the landscape of reverse functional genetics in insects, and has unveiled many gene functions, from development to reproduction, including behavior, coloration, resistance to biological stress, polyphenism, and many others (Belles, 2010).
miRNAs
miRNAs are endogenous, ca. 22-nucleotide, single-strand, non-coding RNAs that regulate gene expression on the post-transcriptional level through base-pairing between the seed sequence of the miRNA and its complementary seed match sequence that is present in the 3′ untranslated region (UTR) of the target mRNA.
The first miRNA, lin-4, was discovered in a screen for genes required for post-embryonic development in the nematode C. elegans (Lee et al., 1993; Ambros and Horvitz, 1984). The identification of the lin-4 locus and its regulatory mechanism through the 3′ UTR of lin-14 mRNA was an interesting finding, although at that time it was almost considered to be a genetic oddity. However, the discovery of another miRNA, let-7, initially in C. elegans (Reinhart et al., 2000) and later in various bila-terian species (Pasquinelli et al., 2000), confirmed that, in the case of lin-4 and lin-14, it was not an oddity at all, but rather a new and fundamental layer of the mechanisms regulating gene expression (Lai et al., 2003; Neilson and Sharp, 2008).
The following sections will deal exclusively with miRNAs.
Biogenesis of miRNAs
miRNAs undergo molecular processing before becoming mature and ready to play their functional role. The pathway of miRNA biogenesis has many commonalities with that of siRNAs, but it is distinct in a number of ways (Figure 2). miRNAs are first transcribed as part of a longer primary transcript (pri-miRNA), which folds, forming hairpin structures that correspond to miRNA precursors (pre-miRNAs). pri-miRNAs are then processed in the nucleus and transported to the cytoplasm, where they undergo final maturation (Figure 2).
miRNA Processing in the Nucleus
Most miRNA genes are transcribed by RNA polymerase II into pri-miRNA, although in some cases the transcription is mediated by RNA polymerase III (Lee et al., 2004a; Borchert et al., 2006). Usually, pri-miRNAs are several kilobases long, contain local stem-loop structures, and are polyadenylated and capped, as in current mRNAs (Cai et al., 2004; Lee et al., 2004a), although the cap and the poly(A) tail are removed during miRNA processing.
Figure 2 Biogenesis of miRNA and siRNA. miRNA gene is transcribed by RNA Pol II/III into a primary transcript (pri-miRNA) that is processed by Drosha/Pasha and exported to the cytoplasm by Exportin-5. In the cytoplasm, the precursor (pre-miRNA) undergoes the final step of maturation and is cleaved by Dicer-1/Loquacious into an miRNA duplex. After the miRNA duplex unwinds, the mature miRNA is maintained with Argonaute-1 protein (Ago-1) forming a RISC which will be coupled to the target mRNA and will degrade, destabilize, or translationally inhibit it, whereas the miRNA* is released and degraded. On the other hand, siRNA is formed when a long and exogenous double-strand RNA (dsRNA) is cleaved by Dicer-2/R2D2 into an siRNA duplex. Likewise with the miRNA pathway, the siRNA duplex is unwound and single-strand siRNAs are maintain with Argonaute-2 protein (Ago-2) forming a RISC which will recognize the target mRNA and degrade it.
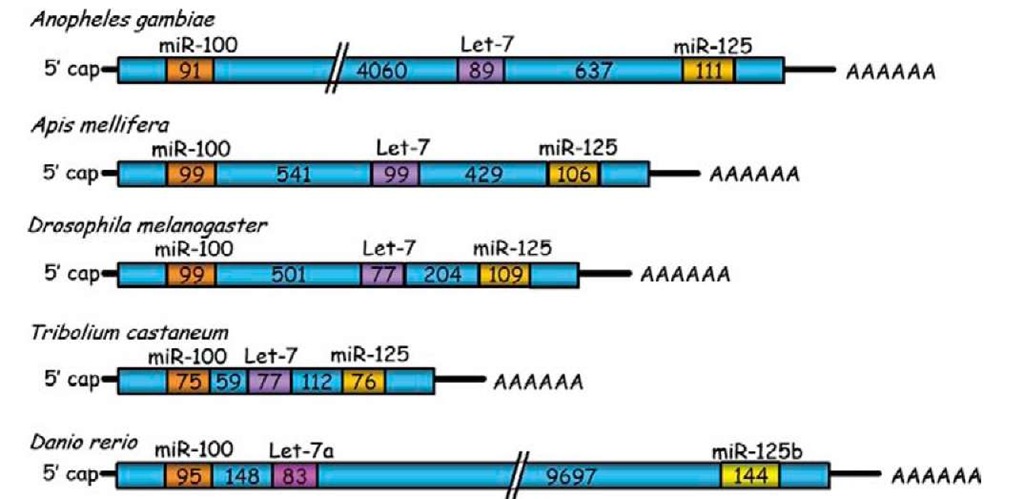
miRNA genes can form clusters in the genome (Behura, 2007), or can be found isolated within an intronic region of protein-coding genes, or in introns and exons of non-coding RNAs (Rodriguez et al., 2004). Moreover, pri-miRNAs can be polycistronic, thus carrying the information of more than one miRNA. In insects, the group of miR-100, let-7, and miR-125 constitutes the best studied example of polycistronic pri-miRNA. The organization of this pri-miRNA is well conserved in many species of insects, and even in vertebrates (Figure 3), although the spacer regions between miRNA precursor sequences can vary considerably in structure and length. For example, the distance between the precursor of miR-100 and that of let-7 varies from ca. 100 bp in T. castaneum to ca. 3.9 kb in Anopheles gambiae, whereas the distance between the precursor of let-7 and that of miR-125 varies within the range of 250-450 bp (Figure 3) (Behura, 2007).
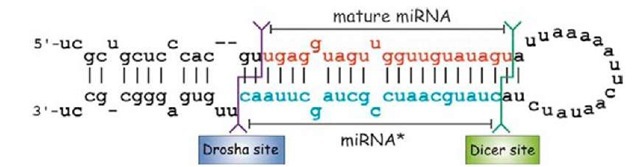
pri-miRNA processing into ca. 70- to 80-nucleo-tide pre-miRNAs takes place exclusively in the nucleus by the action of the microprocessor, a protein complex of ca. 500 kDa, which in D. melanogaster is composed by the RNase III enzyme Drosha and its partner, Pasha (Figure 2) (Denli et al., 2004). In general, insects possess a single pasha gene copy, except in the pea aphid Acyr-thosiphon pisum, where four pasha-like genes have been recently reported (Jaubert-Possamai et al., 2010). Pasha protein (also known as DGCR8 in vertebrates) contains two double-stranded RNA-binding domains; it plays an essential role in miRNA processing by recognizing the substrate pri-miRNA and by determining the precise cleavage site, whereas Drosha actually cleaves the pri-miRNA (Denli et al, 2004). The two RNase domains of Drosha cleave the 5′ and 3′ arms of the pri-miRNA 11 base-pairs away from the single-stranded RNA/double-stranded RNA junction at the base of the hairpin stem (Figure 4) (Han et al., 2004); of note, a single nucleotide variation in an miRNA precursor stem can block Drosha processing (Duan et al., 2007).
Generally, cleavage of the pri-miRNA by Drosha occurs in an unspliced intronic region before mRNA splicing catalysis (Kim and Kim, 2007). However, some miRNA genes are located within intronic regions which themselves form a hairpin structure. In this special case, the action of Drosha is bypassed during pri-miRNA processing. After splicing of its host mRNA, the miRNA is released from the intron, then exported from the nucleus to the cytoplasm, and is finally cleaved by Dicer (see below). This class of miRNA, called mirtrons, has been described in flies, nematodes, and mammals (Berezikov et al, 2007; Okamura et al., 2007; Ruby et al, 2007a).
Figure 3 Organization of the primary transcript of miR-100, let-7, and miR-125 cluster in different insect species and the zebrafish. Numbers inside the boxes correspond to the length in base-pairs.
Figure 4 Precursor of miRNA let-7. In one of the arms of the stem-loop of the hairpin the mature miRNA sequence resides (in red); in another is the miRNA* sequence (in blue). The sites of cleavage for both enzymes Drosha and Dicer-1 are shown as purple and green lines, respectively.
Pre-miRNA Transport from the Nucleus to the Cytoplasm
Once the pri-miRNA is processed in the nucleus, the resulting pre-miRNAs are exported to the cytoplasm by Exportin5 (EXP5) (Figure 2), which is a member of the nuclear transport receptor family, in complex with the cofactor Ran-GTP (Yi et al, 2003; Kim, 2004). EXP5 can recognize double-stranded RNA stems longer than 14 base-pairs along with a short 3′ overhang (1-8 nucleotides), which ensures the export of only those pre-miRNAs correctly processed (Lund et al., 2004). Using RNAi, a number of authors have demonstrated the role of EXP5 in the nucleocytoplasmic transport of pre-miRNA. Knockdown of EXP5 mRNA decreases the levels of mature miRNAs, but does not lead to an increase of pre-miRNA levels in the nucleus, which suggests that protecting pre-miRNA from digestion in the nucleus is another important role of EXP5 (Yi et al., 2003; Lund et al., 2004).
miRNA Maturation by Dicer
In the cytoplasm, pre-miRNAs are cleaved by Dicer near the terminal loop (Figure 4), thus resulting in the release of a ca. 22-nucleotide miRNA duplex with two nucleo-tides protruding as overhangs at each 3′-end. Dicer is an ATP-dependent multidomain enzyme of the RNase family involved in the cleavage of small double-stranded RNAs. It was identified for the first time in D. melanogaster (Bernstein et al., 2001), and two Dicer homologs were later found in this fly, Dicer-1 and Dicer-2, which, in general, are involved in the miRNA and siRNA pathways, respectively (Lee et al., 2004b). RNAi of Dicer-1 in the last instar nymph of the cockroach B. germanica inhibits the formation of mature miRNAs and impairs the meta-morphic process (Gomez-Orte and Belles, 2009), which confirms not only the role of Dicer-1 in miRNA biogenesis in a phyllogenetically basal insect, but also that of miRNAs in hemimetabolan metamorphosis (see below).
D. melanogaster Dicer-1 interacts with the protein Loquacious (also known as R3D1), which contains three double-stranded RNA-binding domains for pre-miRNA processing. Depletion of Loquacious results in pre-miRNA accumulation in Drosophila S2 cells, and immuno-affinity purification experiments revealed that Loquacious locates in a functional pre-miRNA processing complex along with Dicer-1, and stimulates the specific pre-miRNA processing activity (Saito et al., 2005). Of note, both arms of the pre-miRNA stem loop structures are imperfectly paired, containing G : U wobble pairs and single nucleotide insertions (Figure 4). The imperfect base-pairing causes differences in thermodynamic properties and makes one strand of the duplex less stably paired at its 5′ end. Generally, the strand with the lowest thermodynamic stability becomes the mature miRNA (guide strand), whereas the other strand (miRNA* or passenger strand) is degraded.
Regulation of miRNA Biogenesis and Stability
As a general principle, given that most of the miRNA genes are transcribed by RNA Polymerase II, the usual transcription factors associated with Pol II will influence their transcriptional control. A more specific modality of miRNA regulation is the process known as editing, which consists in a post-transcriptional change of RNA sequences caused by deamination of adenosine (A) to inosine (I), thus resulting in alterations in the base-pairing of the transcript. pri-miRNA transcripts modified by ADAR (adenosine deaminase acting on RNAs) have their biogenesis altered in downstream steps. These modifications of the pri-miRNA sequences may block the cleavages by Drosha and Dicer during miRNA maturation; moreover, edited mature miR-NAs can recognize other target mRNAs (Kawahara et al., 2007). Therefore, editing is a remarkable regulator of biogenesis, and in addition increases miRNA structural diversity and further extends the diversity of miRNA targets.
Regulation of the miRNA biogenesis pathway also involves feedback mechanisms, like the interplay of Drosha and Pasha, which regulate each other in a circuit of negative feedback. Drosha acts by cleaving a hairpin located in the 5′ UTR of Pasha mRNA. Excess of Dro-sha decreases Pasha mRNA levels, whereas a reduction of Drosha elicits the reverse effect (Kadener et al., 2009b). Another example of feedback regulation involves human Dicer and the miRNA let-7, wherein Dicer is targeted by let-7 in sites within its coding region (Forman et al., 2008), or the reciprocal regulation showed by let-7 and the RNA-binding protein Lin-28, where let-7 suppresses Lin-28 protein synthesis whereas Lin-28 blocks let-7 maturation. Lin-28 is capable of blocking the cleavages mediated by both Drosha and Dicer; indeed, recombi-nant Lin-28 can block pri-miRNA processing, whereas knockdown of Lin-28 facilitates the expression of mature let-7 (Viswanathan et al., 2008). Lin-28 acts by inducing uridylation of the let-7 precursor at its 3′ end, which elicits the degradation of the uridylated pre-let-7 because Dicer fails to process hairpin RNA structures with long 3′ extensions (Heo et al., 2008).
Unlike 3′ uridylation, 3′ adenylation may have a stabilizing effect on miRNAs, at least in mammals. For example, a variant of miR-122 possesses a 3′-terminal adenosine that is added by cytoplasmic poly(A) polymerase GLD-2 after unwinding of the miR-122/miR-122* duplex, and this 3′ adenylation appears to prevent shortening, thus stabilizing the miRNA (Katoh et al., 2009). Apparently, adenylation and uridylation are two competing processes; it is interesting, in this sense, that addition of adenine residues in some small RNAs can prevent urydilation (Chen et al., 2000).