1. Introduction
An exciting trend in molecular biology involves the use of systematic genomic, proteomic, and metabolomic technologies to construct large-scale models of biological systems. These endeavors, collectively known as systems biology (Ideker etal., 2001a; Kitano, 2002), establish a paradigm by which to systematically interrogate, model, and iteratively refine our knowledge of the cell. While the field of systems science has existed for some time (Ashby, 1958; Bertalanffy, 1973), systems approaches have recently generated a great deal of excitement in biology due to a host of new experimental technologies that are high throughput, quantitative, and large scale. Owing to the time and expense associated with large-scale measurements as well as the enormous amounts of data produced, principled strategies will be indispensable for using these data to construct biological models.
2. Large-scale experimental methods
The first technology to revolutionize modern biology was automated DNA sequencing (Hood etal., 1987), which was instrumental in defining the list of 30 000 genes in the human genome (Lander etal., 2001; Venter etal., 2001). More recently, DNA microarrays have enabled simultaneous measurement of all gene states to reveal which genes are expressed (i.e., turned on vs. off) in a particular cell type or biological condition (Slonim, 2002). Other molecular states, such as changes in protein levels (Gygi et al., 1999), phosphorylation states (Zhou et al., 2001), and metabolite concentrations (Griffin et al., 2001), can be quantified with mass spectrometry, nuclear magnetic resonance, and other advanced technologies. A final cellular state measurement that is gaining in importance is the genomic phenotyping experiment (Begley etal., 2002), also called parallel phenotypic analysis (Deutschbauer etal., 2002). In this type of experiment, a library of gene knockouts is screened to identify which genes are essential for a particular phenotype. In single-celled organisms, the phenotype associated with each gene is typically the growth rate, but it can really be any measure of the phenotypic consequences of perturbing a gene.
Of the approaches for characterizing cellular states, measurements made by DNA microarrays are currently the most comprehensive (every mRNA species is detected); high throughput (a single technician can assay multiple conditions per week); well characterized (experimental error is appreciable, but understood); and cost-effective (whole-genome microarrays are purchased commercially for US $50 to $1000, depending on the organism). However, continued advances in protein labeling and separation technology are making measurement of protein abundance and phosphorylation state almost as feasible, with the primary barrier being the expense and expertise required to set up and manage a mass spectrometry facility. Measurement of metabolite concentrations, an endeavor otherwise known as metabonomics (Nicholson et al., 2002), is currently limited not by detection (thousands of peaks, each representing a different molecular species, are found in a typical NMR spectrum) but by identification (matching each peak with a chemical structure is difficult). Clearly, measuring changes in cellular state at the protein and metabolic levels will be crucial if we are to gain insight into not only regulatory pathways but also those pertaining to the cell’s signaling and metabolic circuitry.
Equally as exciting, another set of recent technological advances have enabled us to characterize DNA and protein interaction networks. Several methods are available for measuring protein-protein interactions at large scale – two of the most popular being the yeast two-hybrid system (Uetz et al., 2000b; Fields and Song, 1989) and protein co-immunoprecipitation (colP) followed by mass spectrometry (Gavin et al., 2002; Ho et al., 2002a). Protein-DNA interactions, which commonly occur between transcription factors and their DNA binding sites, constitute another interaction type that can now be measured at high throughput using the technique of Chromatin ImmunoPrecipitation followed by promoter microarray chip analysis (ChlP-chip) (Iyer et al., 2001; Ren et al., 2000). Large protein-protein or protein-DNA interaction data sets are now available for a variety of species including Saccharomyces cerevisiae (Uetz et al., 2000a; Lee et al., 2002; Ito et al., 2001; Ho etal., 2002b; Gavin et al., 2002), Helicobacter pylori (Rain et al., 2001), Drosophila melanogaster (Giot etal., 2003), and Caenorhabditis elegans (Walhout et al., 2000; Li et al., 2004). Additional types of molecular interactions, such as those between proteins and small molecules (carbohydrates, lipids, drugs, hormones, and other metabolites), are difficult to measure at large scale, although protein array technology (MacBeath and Schreiber, 2000; Zhu et al., 2001; Haab et al., 2001) might enable high-throughput measurement of protein-small molecule interactions in the near future. A current drawback of high-throughput interaction measurements is a potentially high error rate (Deane etal., 2002). An emerging approach for addressing this problem is to construct models that integrate several complementary data sets together (e.g., two-hybrid interactions with coIP data or gene expression profiles) to reinforce the common signal (Bar-Joseph et al., 2003; Hanisch etal., 2002; Ideker etal., 2002; Jansen etal., 2002; Yeger-Lotem and Margalit, 2003; Ge etal., 2001).
3. Modeling and systems analysis
The enormous amount of data arising from high-throughput biology provides a more complete picture of cellular function than ever before. However, new data sets are being generated at a rate that far outpaces our ability to analyze and interpret the results – a disparity that has thus far limited the impact of these data on basic biomedical research. Eliminating this disparity therefore presents a number of grand challenges to computational researchers: how to best associate high-level information about proteins and protein interactions with functional roles; how to enrich the true biological signal in noisy data; and, most importantly, how to organize global measurements at different levels into full-fledged models of cellular signaling and regulatory machinery.
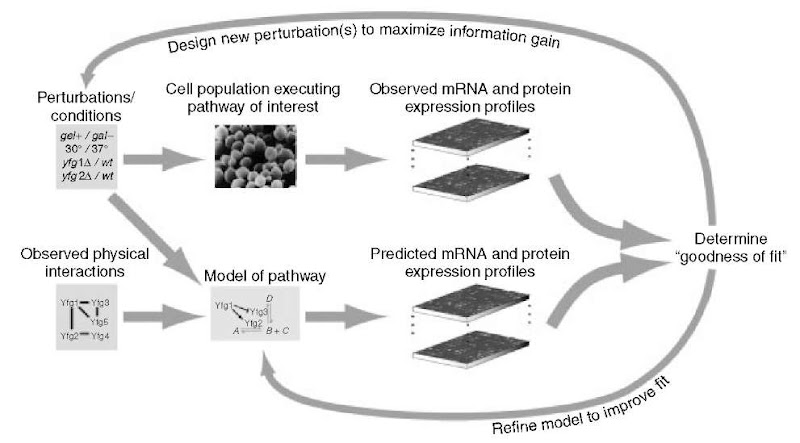
Systems biology attempts to address these goals by integrating the various levels of global measurements together and with a mathematical model of a biological system or pathway of interest. Although these model-driven approaches may differ in the particulars of implementation, all follow a fundamental framework involving the following four distinct steps (Figure 1):
1. Define the System Components. Discover all of the genes in the genome as well as the particular molecules and molecular interactions that constitute the pathway of interest. If possible, define an initial model of how these molecular components and interactions relate to govern pathway function.
2. Perturb the System. Perturb each pathway component through a series of genetic or environmental manipulations. Detect and quantify the corresponding global cellular response to each perturbation, using genomic, proteomic, and/or metabolomic technologies.
Figure 1 A systems approach to biology.
3. Model Reconciliation. Integrate the observed responses with the current pathway model and with the global networks of protein-protein interactions, protein-DNA interactions, and biochemical reactions.
4. Model Verification/Expansion. Formulate new hypotheses to explain observations that are not predicted by the model. Design additional perturbation experiments to test these and iteratively repeat steps (2), (3), and (4).
Steps (1) and (2) are focused on biological discovery through construction of a library of potential components, interactions, and system responses. Steps (3) and (4) are driven by a set of hypotheses encoded by the computational models. Systems approaches following this general framework have been used most actively to interrogate pathways in model organisms, including yeast (Forster et al., 2003; King etal., 2004; Ideker etal., 2001b; Bar-Joseph etal., 2003), Escherichia coli (Gardner et al., 2003), Halobacterium (Baliga et al., 2002), and sea urchin (Davidson et al., 2002). These works provide a roadmap of how to model cellular processes through large-scale measurement and integration of biological data.
Central to this systems approach are computer-aided models for understanding and interrogating complex cellular processes. These models promise to revolutionize biology and medicine by providing a comprehensive blueprint of normal and diseased cell functions and by allowing researchers to simulate the effects of drugs on cells long before they are tested in humans. Several strategies have been developed for integrating gene expression profiles with other large-scale data to formulate models of regulatory networks, including Bayesian learning, neural nets, process algebra, and systems of differential equations [for a review, see van Someren (van Someren et al., 2002)]. Computer-aided approaches for experimental design are equally as important – that is, designing perturbations to most effectively and efficiently verify and expand the models in step (4) above (Tong and Koller, 2001; King et al., 2004; Ideker et al., 2000).
With the recent appearance of large networks of protein-protein and protein-DNA interactions as a new type of measurement, systems researchers are trying to identify the specific network structures and network “design principles” that have been most favored by evolution. These efforts have shown that biological networks encode modular functional units (Rives and Galitski, 2003; Ravasz et al., 2002) that have likely evolved to be robust to perturbation (Jeong etal., 2001). Moreover, these modules often contain recognizable configurations such as the feedback and feed-forward loops that are also prevalent in electronic circuitry and other types of man-made systems (Milo etal., 2002). Several other groups have proposed methods for constructing regulatory models of the cell, using molecular interaction networks as the central framework (Bar-Joseph et al., 2003; Kelley et al., 2003; Hanisch et al., 2002; Ideker et al., 2002; Jansen et al., 2002; Yeger-Lotem and Margalit, 2003; Ge et al., 2001). The key idea is that, by identifying which parts of the molecular interaction network correlate most strongly with other biological evidences such as gene expression profiles or genomic phenotypes, it will be possible to organize the network into circuit modules representing the repertoire of distinct functional processes in the cell. Several approaches are also available for identification of metabolic pathways or protein complexes that have been conserved over evolution (Kelley et al., 2003; Forst and Schulten, 2001; Dan-dekar et al., 1999). Evolutionarily conserved pathways allow interpretation of the network of a poorly understood organism based on its similarity to that of a well-known species. These tools also have application to infectious disease, for example, by targeting drugs to pathways that are present in a pathogenic organism but absent from its human host.
4. Current and future challenges
Systems biology now faces several major challenges. First, given a first-generation toolbox of modeling and systems approaches, an immediate next step is to leverage these tools to elucidate disease pathways. However, although systems approaches have been successfully applied to species such as S. cerevisiae (Gavin et al., 2002; Ho et al., 2002a; Habeler et al., 2002; Gygi et al., 1999; Haab et al., 2001; Kumar et al., 2002; Lee et al., 2002; Tong et al., 2001; Uetz et al., 2000b), similar studies in medically relevant organisms such as human, mouse, and rat have remained largely out of reach. This disparity is due to the increased complexity of mammalian systems; technical and ethical problems in subjecting them to perturbation; and a relative lack of experimental data on protein-protein, protein-DNA, or other molecular interactions for human. Encouragingly, ongoing data production efforts in human cell lines (e.g., the Alliance for Cell Signaling; http://www.afcs.org) and emerging technologies such as RNAi gene knockdowns (Dykxhoorn et al., 2003) promise to address many of these difficulties in the near future. In the meantime, yeast remains an attractive alternative for studying basic biological pathways that influence pathogenesis and genetic disorders.
A second challenge is that almost all previous systems biology studies have been strongly reliant on a preexisting model of the pathway of interest. For instance, in our case study of the galactose-utilization pathway (Ideker et al., 2001b), the initial components and interactions of the model were drawn directly from review articles and the primary literature. An initial literature-based model was also indispensable for the now classic explorations of bacterial chemotaxis (Barkai and Leibler, 1997) and infection by phage lambda (Arkin et al., 1998; McAdams and Shapiro, 1995). These studies significantly expanded our biological knowledge, but if systems approaches continue to be successful, they will quickly exhaust the few well-studied biological systems that are available. Therefore, systems biology will only become a sustainable paradigm if it can generate new models in the absence of extensive prior knowledge.
If these challenges can be met, it is likely that systems and modeling approaches will have substantial payoffs in basic medicine as well as the pharmaceutical industry. In this regard, it is revealing that outside of biotechnology, many sectors of manufacturing already depend heavily on computer simulation and modeling for product development. Using computer-aided design (CAD) tools, digital circuit manufacturers explore the wiring of transistors and other components on the silicon wafer, just as automotive engineers estimate how many miles per gallon is to be expected from the next-generation sedan long before it is built on the assembly line. Biology will undoubtedly also benefit from these “classical” engineering approaches. Given that more than six out of every seven drugs that undergo human testing ultimately fail because of unanticipated side effects, systems modeling may act as a much-needed filter between high-throughput screening for drug candidates and the time-consuming and costly follow up of human trials.
5. Perspective
The field of systems biology still faces many challenges but also holds much promise. By increasing our basic repertoire of experimental strategies and modeling approaches, systems biology provides the starting point for advances in many facets of biotechnology, not least of which is an enhanced ability to appropriately target therapeutics in diseased cells. Thus, we can move one step closer to the day when systems modeling techniques will have widespread influence on basic biological research and replace high-throughput screening as a de facto standard in drug development.