Virusoids are a particular class of plant satellite RNA (Fig. 1). Satellite RNAs are always functionally dependent on specific helper viruses and are encapsidated by the coat protein of these helper viruses. They can be regarded as molecular parasites and in most cases reduce the titer and attenuate the symptom expression of their supporting viruses. Satellite RNAs do not have extensive sequence similarity with the RNA of the helper virus.
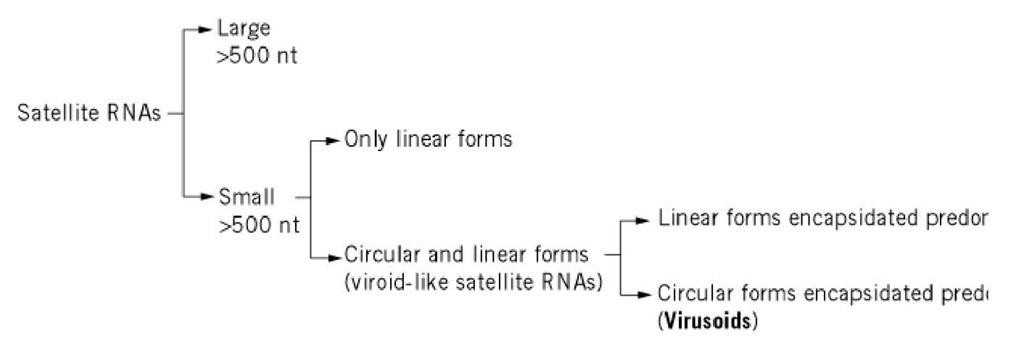
Figure 1. Classification of satellite RNAs from plants.
Figure 1 shows the groups into which plant satellite RNAs can be subdivided. Large satellite RNAs can encode proteins, whereas small satellite RNAs do not. Viroid-like satellite RNAs (Table 1) are single-stranded molecules present in infected tissue in both circular and linear forms, are structurally similar to viroids, and most likely replicate by the same type of rolling circle mechanism. A property that is unique to the viroid-like satellite RNAs of sobemoviruses is that the circular form is the predominant RNA encapsidated by the coat protein of the helper virus, and on this basis the term virusoid has been proposed for them; in contrast, the linear form is predominantly encapsidated in viroid-like satellite RNAs of nepo- and luteoviruses (1). It is also worth noting that at least one plant viroid-like RNA has been found in association with an homologous DNA counterpart forming what has been called a retroviroid-like element (2).
Table 1. Virusoids and Other Viroid-like Satellite RNAs with Their Abbreviations, Genomic Accession Numbers of Typical Sequence Variants, Sizes, and Group to Which They Belong
|
Viroid-like Satellite RNA |
Abbreviation |
Accession |
Size (nt) |
Group |
|
Lucerne transient streak virusoid |
vLTSV |
X01984 |
322324 |
Sobemovirus |
|
Solanum nodiflorum mottle virusoid |
vSNMoV |
J02386 |
377 |
Sobemovirus |
|
Subterranean clover mottle virusoid |
vSCMoV |
M33000 |
322, 388 |
Sobemovirus |
|
Velvet tobacco mottle virusoid |
vVTMoV |
J02439 |
365366 |
Sobemovirus |
|
Rice yellow mottle virusoid |
vRYMV |
AF039909 |
220 |
Sobemovirus |
|
Cereal yellow dwarf virus RPV satellite RNA |
SCYDV-RPV RNA |
M63666 |
322 |
Luteovirus |
|
Tobacco ringspot virus satellite RNA |
sTRSV RNA |
M14879 |
359360 |
Nepovirus |
|
Arabis mosaic virus satellite RNA |
sArMV RNA |
M21212 |
300 |
Nepovirus |
|
Chicory yellow mottle virus satellite RNA |
SCYMoV RNA |
D00721 |
457 |
Nepovirus |
All known viroid-like satellite RNAs contain ribozyme (catalytic RNAs) domains in one or in both polarity strands. The ribozymes of virusoids and sCYDV-RPV RNA are of the hammerhead type and can be adopted by either both polarity strands (vLTSV and sCYDV-RPV RNA) or only their plus polarity strands (vSNMoV, vSCMoV, vVTMoV, and vRYMV). Viroid-like satellite RNAs from nepoviruses can form hammerhead and hairpin (or paperclip) ribozymes in their plus and minus polarity strands, respectively. These ribozymes most probably play a role in replication: hairpin structures mediate both RNA self-cleavage and self-ligation (3), whereas hammerhead structures mediate self-cleavage only (4-6).
It has been proposed that viroid-like satellite RNAs have a common phylogenetic origin with viroids and that they could be molecular fossils of the RNA world (7). The RNA of human hepatitis delta virus shares some properties with viroid-like satellite RNAs, including circular structure, functional dependence on a helper virus, and presence of ribozymes in both polarity strands. One key question to be addressed is the characterization of the determinants responsible for the specificity of the interaction between each viroid-like satellite RNA and its helper virus.