Minicells are small spherical cells that are formed in certain mutants of rod-shaped bacteria. In these mutant strains, the cell division septum is aberrantly placed adjacent to the cell pole instead of at its normal midcell site. The polar septation events give rise to spherical cells (minicells) that lack the bacterial chromosome but otherwise appear normal. The minicells are capable of protein synthesis and normal metabolic functions but are incapable of undergoing further rounds of cell division. The first description of a mutation that led to the minicell phenotype in Escherichia coli was made by Adler et al. (1). Minicell formation has since been described in a number of other Gram-positive and Gram-negative species (2-5).
The formation of polar septa, and the concomitant production of minicells, results from the use of potential division sites that are located at the cell poles. The polar division sites are probably derived from division sites that had been located at midcell during a preceding division event. In this view, components that had been located at the midcell division site are retained at the new poles of the daughter cells after septation is completed, and these sites are still capable of supporting additional septation events (6). It has also been suggested that the polar sites are induced by changes in the organization or intracellular localization of the bacterial chromosomes, rather than being residues of previous division events (7, 8).
The potential division sites at the cell poles are not used to support septum formation because they are suppressed by the gene products of the min genetic locus, as shown by studies of E. coli and Bacillus subtilis. The minicell phenotype occurs when the suppression of polar division events is lost as a result of mutations or experimental manipulations that interfere with function or expression of the min genes (9). The minicell phenotype can also be induced by procedures that increase the expression of the cell division gene ftsZ, thereby increasing the frequency of septum formation at all sites, both the aberrant sites at the poles and the normal division site at midcell (10). The minicell phenotype is also associated with certain mutations or physiological manipulations that perturb DNA structure or replication pattern (8, 11, 12).
Minicelling populations include two types of cell, spherical minicells and rod-shaped cells that are longer than the cells of wild type populations. The lengths of the rod-shaped cells range approximately from one to six times normal cell length. It has been suggested that this occurs because in each division cycle the cell has only enough "division potential" to support a single septation event (one "quantum" of division potential) (6). In this view, if the polar sites are available because of a min mutation, septation can occur either at a polar site or at the normal midcell site; septation at a polar site leads to one minicell and one cell that is almost two cell lengths long. Because a similarly random choice between polar and central division occurs in each generation, a portion of the cells end up being several cell lengths long before a central division occurs. The "division potential" that is postulated in this quantal model of cell division could correspond to the cellular content of FtsZ, because FtsZ appears to be the rate-limiting protein in the division process (see Cell division). The quantal hypothesis has recently been questioned on the basis of examination of the division pedigrees of individual cells (8) but still remains an attractive explanation of the known facts.
Minicell mutants have been used primarily for two purposes: (1) to better understand the mechanism of cell division, with emphasis on the mechanism used by the cell to identify the correct location for the division site; and (2) as factories for the synthesis and characterization of proteins.
1. E. coli min Genes
Most of what is known about the normal regulation of polar division sites has come from studies of min mutants of E. coli (13). The min genetic locus defines an operon that includes three genes, minC, minD, and minE (9). This gene cluster is sometimes called the minB locus. The term minB is a historical anachronism, resulting from the original misconception that mutations in two genetic loci, called minA and minB, were required to produce the minicell phenotype. It was later discovered that the original identification of minA was an error and that the minicell phenotype resulted from a single mutation in minB (ie, in the minCDE gene cluster) (14, 15). This mutation later was shown to reside in minD (16).
The minCDE gene products act in concert to prevent septation at the cell poles. This is accomplished by the coordinated action of a nonspecific division inhibitor, whose activity requires the expression of minC and minD, and a topological specificity factor (MinE) that restricts the action of the division inhibitor to the polar division sites (Fig. 1). As a result, minicell formation is prevented, whereas septation at the midcell site occurs normally. Consistent with this model, loss of either MinC or MinD leads to septation at the polar sites, with production of the minicell phenotype.
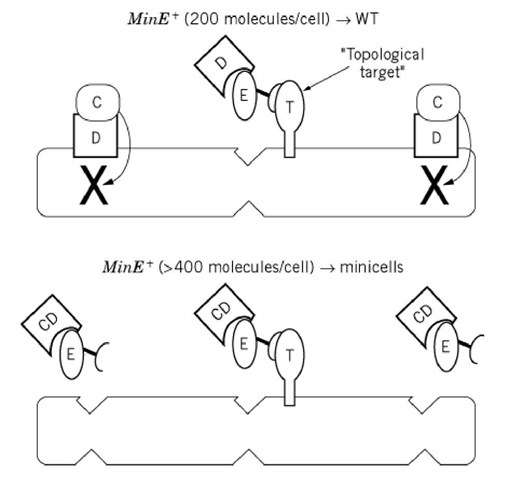
Figure 1. Current model for function of min gene products.
MinC and MinD act in concert to produce a nonspecific inhibitor of septation, as shown by the observation that expression of minC and minD in the absence of minE leads to the formation of long nonseptate filaments. In the MinCD filaments, septation is prevented at the cell poles, as well as at the normal midcell site. Although at normal levels of expression MinC and MinD are both required for the division block, MinC is believed to be the actual division inhibitor protein. This conclusion is based on the observation that high levels of expression of MinC lead to the formation of nonseptate filaments even in the absence of MinD (17). MinD is believed to function as an activator of the latent or low-level division inhibition activity of MinC. Studies using the yeast two-hybrid system have indicated that MinC and MinD can interact (18).
The MinCD division inhibitor blocks septation by interfering with the earliest detectable step in cell division, the formation of the cytokinetic FtsZ ring that appears at the division site prior to septal ingrowth (19) (see Cell division). Consistent with the idea that FtsZ is the MinCD target, the ability of MinCD to block division is suppressed by the overexpression of FtsZ (20, 21).
MinC also plays an essential role in a second division inhibition system, the minC-dicB system. In this system, DicB plays the role that MinD plays in the minC-minD system. The dicB gene is believed to be part of a cryptic prophage in the E. coli chromosome (22). Its expression is normally tightly repressed, and there are no known physiological mechanisms to relieve this repression. However, when dicB is induced by placing it under control of a regulatable promoter, such as P^ (see Lac Operon), division is blocked completely, leading to formation of long nonseptate filaments (22). The DicB-mediated division block requires the presence of MinC but is independent of MinD (20). Taken together with the observation that high concentrations of MinC can block division in the absence of MinD or DicB (17), this has led to the conclusion that DicB and MinD can act as independent cofactors or activators of the MinC-mediated division inhibition reaction. It is not known how DicB or MinD function to activate the division inhibition systems.
DicB and MinD have no apparent sequence similarity, and it is likely that the two proteins use different mechanisms to activate the division inhibition reaction. The MinD protein has ATPase activity, consistent with the presence of a predicted nucleotide-binding motif in the protein sequence (23). Mutational inactivation of this site results in loss of the ability of MinD to activate the MinC-mediated division inhibition reaction. This suggests that nucleotide binding and/or hydrolysis play a role in MinD function. In contrast, the dicB sequence shows no similar nucleotide-binding site. A number of minC mutations have been identified that inactivate the division inhibition activity of the MinCD or MinC-DicB systems. Some of the mutants showed some degree of preferential resistance to either DicB or MinD (24), suggesting that the two proteins may act on different regions of MinC or on different aspects of the division inhibition system. The minC-minD and the minC-dicB division inhibition reactions also can be distinguished by differences in their response to the MinE protein (discussed below).
When minE is expressed coordinately with minCD, the MinCD-induced filamentation is prevented and the wild type division pattern is restored. Thus, MinE prevents the MinCD division inhibitor from working at midcell, without interfering with its ability to prevent minicell formation by blocking septation at the cell poles. This implies that MinE, either directly or indirectly, can distinguish between the new potential division site at midcell and the potential division sites at the cell poles.
The ability of MinE to suppress the activity of the MinC-mediated division inhibition reaction depends on the presence of MinD. Thus, although MinE can prevent MinCD-mediated filamentation, it has no effect on the division block that results from the coexpression of MinC and DicB, nor on the division block that results from high-level expression of MinC alone. Therefore, it is thought that MinD performs two roles. First, it activates the MinC division inhibitor. Second, it mediates the interaction between MinE and the division inhibition system.
To explain the site-specificity of MinE action, a model has been proposed in which MinE has a high affinity for a topological target protein that acts as a MinE docking protein or receptor (Fig. 1) (25). The target protein is presumed to be located at midcell, where it sequesters MinE. As a result, MinE only counteracts the MinCD division inhibitor at the midcell site. This model is consistent with the low abundance of MinE (approximately 200 monomers per cell) and with the observation that a twofold or greater increase in the cellular concentration of MinE leads to minicell formation. According to this model, the increased level of MinE saturates the topological target, leaving the excess MinE free to counteract the MinCD division inhibitor at the polar sites, leading to the observed minicell phenotype. As predicted from this model, the effect of the excess MinE can be counteracted by increasing the level of MinCD, which restores the wild-type division pattern, presumably due to titration of the excess MinE. Thus it is the relative concentrations of MinC, MinD, and MinE that are important, rather than the absolute concentration of the individual components.
In a variation of this model, the topological target is located at the cell poles instead of midcell (26). This would lead to the sequestration of MinE at the polar sites, where it would play a positive role, activating the division inhibitor at the cell poles. The evidence available at this time favors the first model in which the MinE target is at midcell; but until the locations of the proteins are directly determined, both models are tenable.
Two functions are defined for MinE. First, it is an antagonist of the MinCD division inhibitor, as shown by its ability to prevent the formation of nonseptate MinCD filaments. Second, it is a topological specificity factor, limiting the action of the division inhibitor to the potential division sites at the cell poles. Genetic deletion analysis has shown that the anti-MinCD function resides entirely within the 22-residue N-terminal domain of MinE, MinE . Thus, expression of MinE prevents MinCD-induced filamentation (25). However, the MinE domain lacks the topological specificity function of MinE, because expression of MinE in the presence of MinCD leads to a minicell phenotype instead of a wild-type division pattern. In contrast, deletion analysis has shown that the topological specificity function of MinE, as defined by its ability to prevent minicelling, requires a domain that extends close to the carboxy-terminus of the protein (25, 26).
MinE is probably an oligomeric protein, as shown by gel filtration studies using MalE-MinE chimeric proteins. These studies, and studies using the yeast two-hybrid system, have indicated that the domain required for oligomerization lies within the carboxy-terminal half of the protein (26).
2. min Genes in B. subtilis
Mutations in two B. subtilis genetic loci, divIVA and divIVB, have been shown to lead to minicell phenotypes (2). The divIVB locus includes homologues of the E. coli minC and minD genes (27-29). The homology of the predicted amino acid sequences is strong for MinD (43.7% identity and 83.3% similarity between the E. coli and B. subtilis proteins) and less so for MinC (17.7% identity and 26.1% similarity). Similar to the results in the E. coli system, insertional disruption of the B. subtilis minD or minC gene leads to a strong minicell phenotype. However, in contrast to E. coli, where minE is located immediately downstream of minD, a minE homologue has not been identified in B. subtilis.
It is not known whether the B. subtilis MinC and MinD proteins function like their E. coli homologues. Similarity in function is suggested by the observation that minicelling results from mutation of minC or minD in both species, and that expression of the B. subtilis proteins in wild-type E. coli leads to minicelling (27). This suggests that B. subtilis MinC and/or MinD can compete with their E. coli counterparts and implies some similarity in function. It is possible that the Min proteins function similarly in the two species, but that the topological specificity function that is carried out by MinE in E. coli is incorporated into the MinC or MinD protein of B. subtilis, thereby eliminating the need for a separate minE gene. Alternatively, a B. subtilis minE homologue could be located within the adjacent gene cluster that was expressed together with minC and minD when minicelling was induced in B. subtilis, or a B. subtilis minE counterpart could be located elsewhere in the chromosome. This would be consistent, for example, with the observation that overexpression in B. subtilis of the B. subtilis minC and minD genes (together with adjacent genes) leads to a minicell phenotype (27). A similar result has been observed in E. coli when minCDE is overexpressed (30).
Mutations in a second B. subtilis locus, divIVA, also lead to a minicelling phenotype (2, 31). The divIVA gene codes for a 264-residue protein (31). The phenotype resulting from inactivation of divIVA differs from that of DivIVB (minC or minD) minicelling mutants, because the divIVA mutants show a lower frequency of minicelling and also show extensive filamentation.
Overexpression of divIVA leads to a block in cell division, as shown by the formation of long nonseptate filaments (31). The DivIVA-induced division block appears to require a functional MinC protein, since filamentation was not observed when divIVA was overexpressed in a MinC mutant strain. Further evidence for a functional interaction between DivIVA and MinCD comes from the observation that the phenotype of a divIVA-minC double mutant resembles that of a minC single mutant rather than that of the divIVA mutant alone (31). Taken together with the observation that DivIVA-mediated division inhibition requires MinC function, this implies that MinCD acts downstream of DivIVA and is consistent with the view that the divIVA gene product may act by modulating the activity of MinCD. This could occur, for example, if DivIVA acted as an activator of a latent division inhibitor activity of MinC, thereby functionally resembling the E. coli MinD and DicB proteins (discussed above).
3. FtsZ and Minicell Formation
Moderate overexpression of the essential cell division protein FtsZ leads to minicell formation (10). However, in contrast to the phenotype associated with mutation of the min genes, where the population consists of a mixture of spherical minicells and of rod-shaped cells that are longer than wild-type cell length, overexpression of FtsZ leads to a mixture of spherical minicells and of rod-shaped cells that are shorter than the cell length of wild-type cells. This occurs because the increased concentration of FtsZ leads to an increase in the frequency of septation events, with the extra septations occurring at the normally unused polar sites. As a result, minicell formation leads to the generation of one minicell and one cell that is shorter than normal. Presumably, the excess FtsZ overcomes the normal MinCD-mediated suppression of the potential division sites at the cell poles. This is consistent with the observation that overexpression offtsZ suppresses the filamentation that otherwise occurs when the cellular concentration of MinC and MinD is increased above normal levels.
4. Effect of Minicell Mutations on Chromosomal Organization
Several observations suggest a relationship between minicell formation and chromosome location or the state of chromosomal organization. Akerlund et al. (8) have shown that minicell formation occurs when aberrant chromosome replication takes place in cells in which oriC is nonfunctional and a plasmid R1 replication origin is present elsewhere in the chromosome. Minicelling also has been reported to occur when the DNA-binding Fis protein is nonfunctional, and in gyrA and gyrB mutants (11, 12), although in the gyr mutants there is apparently a continuum of cell lengths ranging from "classic" spherical minicells to cells of approximately normal or longer than normal cell lengths. In all of these cases, the nucleoid-free regions at the cell poles appear longer than normal. In addition, the nucleoid-free region at the poles has been reported to be longer than normal in min mutants (7), and it has been reported that minicell mutants produce a low but significant number of anucleate cells of normal cell length (32). An altered degree of supercoiling of some plasmids has also been reported to occur in minicell mutants (7). On the basis of these observations, it has been suggested that the min gene products act on chromosome organization or partition, leading in min~ mutants to a nucleoid-free zone at the cell poles. In this view, polar septation occurs as the default mode wherever a sufficiently large nucleoid-free zone exists; and the Min proteins do not act on old division sites at the poles, but directly on the nucleoid or on the chromosome replication or partition system (7, 8).
5. Synthesis of Plasmid-Encoded Proteins in Minicells
Extrachromosomal genetic elements such as plasmids segregate into minicells, although the bacterial chromosome does not. Therefore, when a minicelling mutant strain is transformed with a native or recombinant plasmid, the minicells contain plasmid DNA and continue to synthesize plasmid-encoded message after the short-lived chromosomally encoded messenger RNA has decayed. Therefore, when the minicell population is labeled with a radioactive amino acid, essentially all of the labeled proteins that are synthesized are encoded by the plasmid DNA. This has been exploited as a generally applicable method to identify the gene products of desired genes by using plasmids that include the genes of interest (33).