Light in the ultraviolet (UV) and visible (vis) range of the electromagnetic spectrum shows an energy that is equivalent to about 150 to 400 kJ/mol. Light with the appropriate energy is used to promote electrons from the ground state to an excited state. The absorption of energy from the incident light as a function of its wavelength is measured in absorption spectroscopy. Molecules with electrons that participate in delocalized aromatic systems often absorb light in the near-UV or visible region.
Absorption spectroscopy is usually performed on solutions of molecules in a transparent solvent. The absorbance of a solute depends linearly on its concentration, and therefore absorption spectroscopy is ideally suited for quantitative measurements. The spectral properties of a molecule depend on the molecular environment and the mobility of its chromophores. Absorption spectroscopy and difference spectroscopy (see Difference Spectroscopy) are therefore well-suited to follow enzyme-catalyzed reactions, ligand binding, and conformational transitions in proteins and nucleic acids. Spectroscopic measurements are very sensitive, nondestructive, and require only small amounts of material for analysis. Spectrophotometers are standard laboratory equipment, and the measurement of absorbance is technically simple.
1. Absorbance of Proteins
In proteins, the peptide bond absorbs light in the range of 180 to 230 nm (which is called the "far-UV" range). The aromatic residues, tyrosine (Tyr), tryptophan (Trp), and phenylalanine (Phe), also absorb light in this region and, in addition, show bands near 260 to 280 nm (in the "near-UV"). Disulfide bonds absorb weakly near 260 nm. Some protein cofactors, such as the heme group, show absorbance in the visible range. When the peptide groups and aromatic residues are part of an asymmetric structure, or when they are immobilized within an asymmetric environment (as in folded proteins), left- and right-handed circularly polarized light is absorbed to different extents. This phenomenon is called circular dichroism (CD).
Spectra of the aromatic amino acids and model proteins are found in (1). In the near-UV, the molar absorbance of a phenylalanine residue is much smaller than that of a tyrosine or a tryptophan, and the spectrum of a protein between 240 and 300 nm is therefore dominated by the contributions from the Tyr and Trp side-chains. Phe residues contribute fine structure ("wiggles") to the spectrum between 250 and 260 nm. The aromatic amino acids do not absorb above 310 nm, and therefore protein absorbance should be zero at wavelengths greater than 310 nm. Proteins without Trp residues do not absorb above 300 nm (see Fig. 1c).
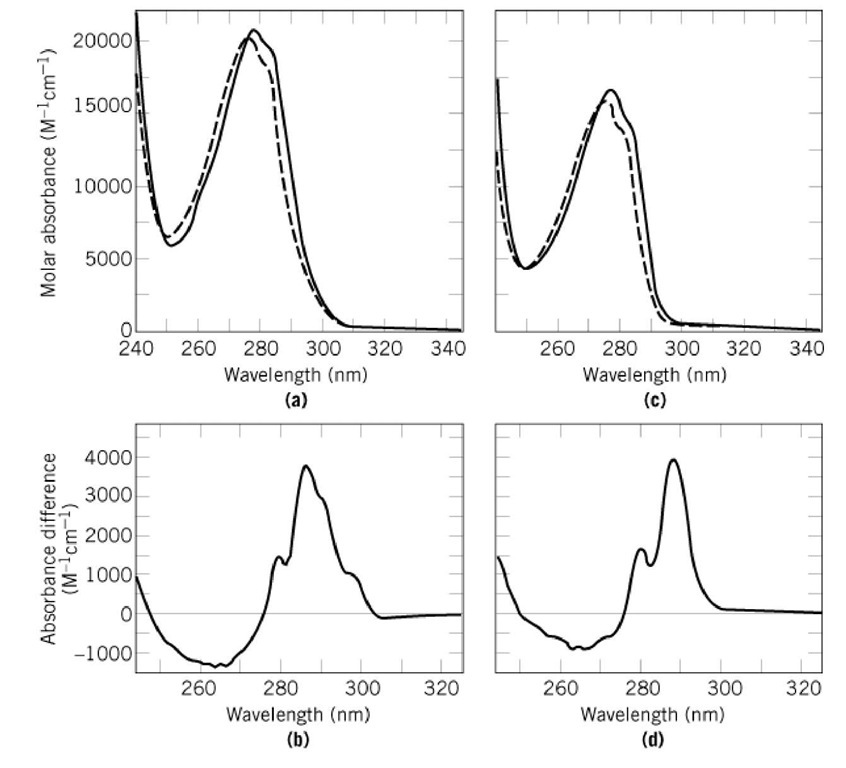
Figure 1. Ultraviolet absorption spectra of (a) the wild-type form and (c) the Trp59Tyr variant of RNase Tr The spectra of the native proteins (in 0.1 M sodium acetate, pH 5.0) are shown by the continuous lines. The spectra of the unfolded proteins (in 6.0 M GdmCl in the same buffer) are shown by broken lines. The difference spectra between the native and unfolded forms are shown in (b) and (d). Spectra of 15 ^M protein were measured at 25°C in 1-cm cuvettes in a double-beam instrument with a band width of 1 nm at 25°C. The spectra of the native and unfolded proteins were recorded successively, stored, and subtracted.
A fraction of the aromatic residues are buried in the hydrophobic core of a native, folded protein molecule (see Protein Structure). When these residues become exposed to the aqueous solvent upon unfolding, their absorption is shifted slightly to shorter wavelengths. This is clearly seen for two forms of ribonuclease T1 (Figs. 1a and 1c). The difference spectra in Figures 1b and d show that the maximal differences in absorbance occur in the 285 to 295 nm region. The difference spectrum for the form without a Trp residue (Fig. 1d) shows a prominent maximum at 287 nm. It is typical for proteins that contain Tyr residues only. The form with a single Trp residue (Fig. 1b) shows additional shoulders between 290 and 300 nm in the difference spectrum. They originate from the single Trp residue of this protein. The differences in the absorption spectra between the native and unfolded state of a protein are generally small, but they can be determined with good accuracy and are extremely useful for monitoring conformational changes of a protein.
Spectroscopic methods are, in general, the methods of choice (i) to investigate changes in the behavior of a protein under different solvent conditions, and (ii) to compare the properties of related molecules, such as homologous or mutated forms of a protein. In addition, they are widely used to measure protein stability and to follow structural transitions such as unfolding and refolding under a variety of conditions. Absorbance changes during fast reactions that occur in the milliseconds range can be followed by using rapid mixing techniques, such as in stopped-flow spectrometry (see Kinetics).
2. Absorbance of Nucleic Acids
Nucleic acids show a strong absorbance in the region of 240 to 275 nm. It originates from the![]()
![]() transitions of the pyrimidine and purine ring systems of the nucleobases. The bases can be protonated, and therefore the spectra of DNA and RNA are sensitive to pH. At neutral pH, the absorption maxima range from 253 nm (for guanosine) to 271 nm (for cytidine), and therefore polymeric DNA and RNA show a broad and strong absorbance near 260 nm.
transitions of the pyrimidine and purine ring systems of the nucleobases. The bases can be protonated, and therefore the spectra of DNA and RNA are sensitive to pH. At neutral pH, the absorption maxima range from 253 nm (for guanosine) to 271 nm (for cytidine), and therefore polymeric DNA and RNA show a broad and strong absorbance near 260 nm.
In native DNA, the bases are stacked in the hydrophobic core of the double helix, and therefore their absorbance is considerably decreased relative to the absorbance of single-stranded DNA, and even more so relative to oligonucleotides. This phenomenon is called hypochromism. It is widely used to follow the melting of DNA double helices.
3. Absorbance to Determine Concentrations
Absorbance measurements are the methods of choice to determine the concentration of proteins or nucleic acids in solution. Spectrophotometers are standard laboratory equipment, and absorbance can be measured quickly and accurately. The absorbance A is related with the intensity of the light before I0 and after I passage through the protein solution by Equation 1, and the absorbance depends linearly on concentration, according to the Lambert-Beer relationship (Eq. 2):
where c is the molar concentration, / the pathlength in cm, and e the molar absorption coefficient. The concentration of a substance in solution can be determined very rapidly and accurately from its absorbance by using Equation 2. The measurement of absorbance values greater than 2 should be avoided, because only 1% of the incident light is transmitted through a solution with an absorbance of 2 (and is quantified by the photomultiplier). The value of e can be determined by a number of experimental techniques (2) or can be calculated for proteins by adding up the contributions of the constituent aromatic amino acids of a protein (2, 3).
The absorbance of nucleic acids does not vary much. The concentrations of nucleic acids in solution are routinely determined from the absorbance at 260 nm. In fact, amounts of nucleic acid are often given as "A260 units." For double stranded DNA, one A^ unit is equivalent to![]() ; for single-stranded DNA, it is equivalent to
; for single-stranded DNA, it is equivalent to![]() ; for single-stranded RNA, it is equivalent to
; for single-stranded RNA, it is equivalent to ![]() All these amounts would produce an A260 of 1 when dissolved in 1 mL and measured in a 1-cm cuvette. Proteins absorb much more weakly than nucleic acids. In a 1:1 mixture of nucleic acids and proteins, the proteins contribute only about 2% to the total absorbance at 260 nm. Consequently, these quantities of contaminating protein hardly affect the concentrations of nucleic acids measured by A260.
All these amounts would produce an A260 of 1 when dissolved in 1 mL and measured in a 1-cm cuvette. Proteins absorb much more weakly than nucleic acids. In a 1:1 mixture of nucleic acids and proteins, the proteins contribute only about 2% to the total absorbance at 260 nm. Consequently, these quantities of contaminating protein hardly affect the concentrations of nucleic acids measured by A260.
4. Absorbance Spectrophotometers
Absorbance is measured by a spectrophotometer. Spectrophotometers consist usually of two light sources: a deuterium lamp, which emits light in the UV region, and a tungsten/halogen lamp for the visible region. After passing through a monochromator (or through optical filters), the light is focused into the cuvette, and the amount of light that passed through the sample is detected by a photomultiplier or photodiode. In diode array spectrophotometers, the sample is illuminated by the full lamp light; after passage through the cuvette, the transmitted light is spectrally decomposed by a prism into the individual components and quantified by an array of diodes, often in intervals of 2 nm. In diode array spectrometers, the entire spectrum is recorded at the same time and not by a time-dependent scan as in conventional instruments. This is of advantage for measuring time-dependent changes at several wavelengths simultaneously (see Kinetics).
5. Buffers for Absorbance Spectroscopy
Good buffers for measuring difference spectra ideally should not absorb light in the wavelength range of the experiment. For work in the near-UV, buffer absorbance should be small above 230 nm, and indeed most of the solvents commonly used in biochemical experiments do not absorb in this spectral region (1). Buffer absorbance is a major problem, however, in the far-UV region below 220 nm, because buffers that contain carboxyl and/or amino groups absorb light in this wavelength range. Buffers with negligible absorbance in the far-UV include phosphate, cacodylate, and borate. Detailed procedures for the measurement of difference spectra are found in Ref. 1.