Introduction
More than half a century ago, Barbara McClintock’s observation of unstable mutations in maize led to the discovery of two mobile genetic elements, Activator (Ac) and Dissociator (Ds) (McClintock, 1948, 1950). Her discovery of these mobile segments of DNA, later named transpos-able elements (TEs), set forth the revolutionary concept of a fluid and dynamic genome. Six decades later, as biology is entering the post-genomic era, there is renewed and rapidly growing appreciation for the tremendous diversity of TEs and their evolutionary impact.
Being mobile, TEs have the ability to replicate and spread in the genome as primarily "selfish" genetic units (Doolittle and Sapienza, 1980; Orgel and Crick, 1980). They tend to occupy significant portions of the eukary-otic genome. For example, at least 46% of the human genome (Lander et al., 2001) and 47% of the yellow fever mosquito genome (Nene et al., 2007) are TE-derived sequences. The relative abundance and diversity of TEs have contributed to the differences in the structure and size of eukaryotic genomes (Kidwell, 2002; Feschotte and Pritham, 2007). TE insertion and recombination are major sources of potentially detrimental mutations, and the host genomes have evolved sophisticated mechanisms to control TE activity (see, for example, Hartl et al., 1997; Malone and Hannon, 2009). The same inser-tional or recombinatory activities by TEs generate a great deal of genetic and genomic plasticity, and provide the raw material for adaptive evolution (Kidwell and Lisch, 2000; Brookfield, 2005; Feschotte and Pritham, 2007; Lin et al., 2007; Cordaux and Batzer, 2009; Gonzalez and Petrov, 2009). For example, TEs have reshaped the human genome by ectopic rearrangements, by creating new genes, and by modifying and shuffling existing genes (Lander et al., 2001; Muotri et al., 2007; Cordaux and Batzer, 2009). In some cases, TEs had been co-opted to perform critical functions in the biology of their host. One well-documented example is the generation of the extensive array of immunoglobulins and T-cell receptors by V(D)J recombination, which is evolved from an ancient transposition system (Gellert, 2002; Fugmann, 2010). Another example is the maintenance of telomeric structures in Drosophila melanogaster by site-specific insertions of two TEs (Pardue and DeBaryshe, 2002; Mason et al., 2008). Therefore, the "selfish" TEs could evolve a wide spectrum of relationships with their hosts, ranging from "junk parasites" to "molecular symbionts" (Brook-field, 1995; Kidwell and Lisch, 2000). Moreover, the apparent arms race between TEs and the host genomes has driven the evolution of the recently discovered piRNA (Piwi-interacting RNA) and endogenous small interfering RNA (endo-siRNA) pathways, which may have had profound impacts on gene regulation and epigenetic silencing (Aravin et al., 2007; Nishida et al., 2007; Pelisson et al., 2007; Yin and Lin, 2007; Brennecke et al., 2008; Chung et al., 2008; Ghildiyal et al., 2008; Klattenhoff et al., 2009; Lau et al., 2009; Lisch, 2009; Malone and Hannon, 2009; Zeh et al., 2009).
The intricate dynamic between TEs and their host genomes is further complicated by the fact that some TEs are capable of crossing species barriers to spread in a new genome. Such a process is referred to as horizontal (or lateral) transfer, which is distinct from the vertical transmission of genetic material from ancestral species/ organisms to their descendants. Horizontal transfer may be an important part of the life cycle of some TEs, and it may contribute to their continued success during evolution (Silva et al., 2004). The recent explosion of genome sequencing projects has provided convenient resources and revealed additional examples and novel insights into horizontal transfer.
From an applied perspective, TEs have been used as tools to genetically manipulate cells/organisms, taking advantage of their ability to integrate cognate DNA in the genome. A well-known example is the transformation system derived from the D. melanogaster P transposable element, which has been instrumental to our understanding of this model organism by providing transformation and mutagenesis tools.In addition, some TEs have been used as genetic markers for mapping and population studies, taking advantage of their dimorphic insertion states (presence and absence of an insertion) and their interspersed distribution in the genome. For example, the human Alu elements have been shown to be useful population genetic markers (Batzer et al., 1994; Batzer and Deininger, 2002; Salem et al., 2003; Ray, 2007). Similar types of markers have been used to trace the explosive speciation of the Cichlid fishes and other vertebrates (Shedlock and Okada, 2000; Terai et al., 2003). TEs have also been used as markers in insects to study incipient spe-ciation, and to map resistance genes (Barnes et al., 2005; Bonin et al., 2008, 2009; Santolamazza et al., 2008).
In this topic, I provide an update to the previous review (Tu, 2005) and focus on recent advances in the study of insect TEs. A brief introduction on TE classification and transposition mechanisms will be followed by sections that descibe the current approaches to studying insect TEs, and sections that highlight the impact and the evolutionary dynamics of TEs in insect genomes. Applications of TEs in genetic and molecular analysis of insects will be discussed towards the end of the topic. Readers may consult recent reviews for details on related topics.
Figure 1 Mechanism of generating target site duplication (TSD). Both sides of the TSD are not part of the TE sequence; they are target sequences duplicated upon a TE insertion. Most TEs create TSDs, although the Helitron DNA transposons and some non-LTR retrotransposons do not.
Classification and Transposition Mechanisms of Eukaryotic Transposable Elements
TEs can be categorized as Class I RNA-mediated or Class II DNA-mediated elements, according to their transposition mechanisms (Finnegan, 1992). The transposition of RNA-mediated TEs involves a reverse transcription step, which generates cDNA from RNA molecules (Eickbush and Malik, 2002). The cDNA molecules are then integrated in the genome, allowing replicative amplification. The transposition of DNA-mediated elements is directly from DNA to DNA, which does not involve an RNA intermediate (Craig, 2002). In most cases, both classes of TEs will create target site duplication (TSD) upon their insertion in the genome (Figure 1). Both DNA-mediated and RNA-mediated elements can be further categorized into different groups. There have been several reviews on different classes of TEs (Deininger and Roy-Engel, 2002; Eickbush and Malik, 2002; Feschotte et al., 2002; Robertson, 2002). All groups of TEs discussed here have been found in various species of insects.
Class I RNA-Mediated TEs
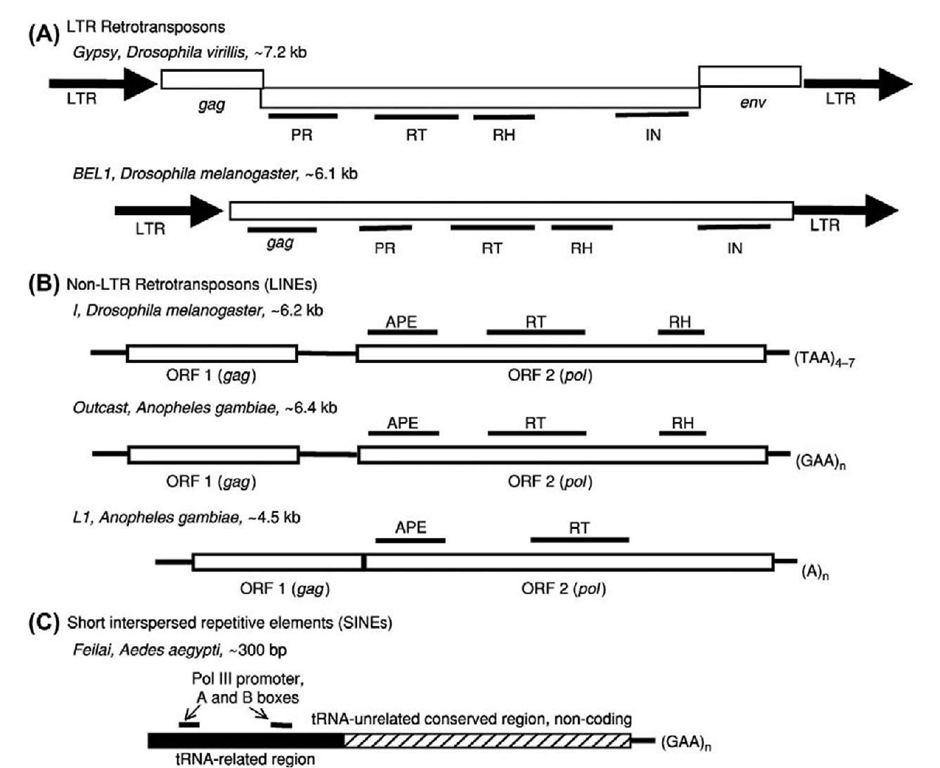
RNA-mediated TEs include long terminal repeat (LTR) retrotransposons, non-LTR retrotransposons, and short interspersed repetitive/nuclear elements (SINEs). Non-LTR retrotransposons are also referred to as retropo-sons or long interspersed repetitive/nuclear elements (LINEs). The structural features of the three groups of RNA-mediated TEs are illustrated in Figure 2, using representatives from different insects. All RNA-mediated TEs produce RNA transcripts that are reverse transcribed into cDNA to be integrated in the genome (Eickbush and Malik, 2002). Detailed mechanisms used by LTR and non-LTR retrotransposons are elegantly described in recent reviews (Eickbush, 2002; Voytas and Boeke, 2002; Eickbush and Jamburuthugoda, 2008).
LTR retrotransposons LTR retrotransposons transpose through a mechanism much like that used by retroviruses. The LTRs in the LTR retrotransposons are generally 200-500 bp long, and are involved in all aspects of their life cycle that include providing promoter sequences and transcription termination signals (Eickbush and Malik, 2002). As shown in Figure 2, LTR retrotransposons encode a pol (polymerase)-like protein that contains reverse transcriptase (RT), ribonuclease H (RNase H), protease (PR), and integrase (IN) domains that are important for their retrotransposition. The RT domain performs the key function ofreverse transcription, and its sequence has been used for phylogenetic classification of LTR retrotransposons into four clades: Tyl/copia; Ty3/gypsy; BEL; and DIRS (Eickbush and Malik, 2002). The IN domain is responsible for inserting the cDNA copy into the host genome. In addition to the pol-like protein, LTR retrotransposons encode an additional protein related to the retroviral gag (group-associated antigene, or group-specific antigen) protein that binds nucleic acids or forms the nucleocapsid shell. Some LTR retrotransposons also have an env (envelope)-like fragment that encodes a transmembrane receptor-binding protein that allows the transmission of retroviruses. Some of the LTR retrotransposons that encode an env protein are in fact retroviruses (Eickbush and Malik, 2002). Some LTR retrotransposons use a tyrosine recombinase instead of the integrase to integrate into the host genome (Eickbush and Jamburuthugoda, 2008).
Non-LTR retrotransposons Non-LTR retrotransposons, or LINEs, or retroposons, are generally 3-8 kilobases long, and have been found in virtually all eukaryotes studied.
Figure 2 Structural characteristics of representative Class I RNA-mediated transposable elements in insects. Representatives are shown from three major groups, long terminal repeat (LTR) retrotransposons (A), non-LTR retrotransposons (B), and short interspersed repetitive elements (SINEs, (C)). The name of each representative element, its host species, and its approximate length are shown as the heading. Open reading frames (ORFs) are shown as open boxes. Env, envelope protein; gag, group-associated antigene, or group-specific antigen; IN, integrase; LINEs, long interspersed repetitive elements; LTR, long terminal repeat; PR, protease; RH, RNase H; RT, reverse transcriptase. The elements are not drawn to scale.
Like the LTR retrotransposons, most non-LTR retrotransposons also have a pol-like protein that includes an RT domain which is essential for their retrotransposition. The RT domain has been used for phylogenetic classification of non-LTR retrotransposons into 17 clades, most of which probably date back to the Precambrian era, approximately 600 million years ago (Eickbush and Malik, 2002; Biedler and Tu, 2003). Some elements also have an RNase H and/or AP endonuclease (APE) domain encoded in the pol-like open reading frame. In addition to the pol-like protein, many non-LTR retrotransposons encode an additional protein related to the retroviral gag protein. Studies of a gag-like protein from L1 retrotransposon in mice show that it acts as a nucleic acid chaperone (Martin and Bushman, 2001). Other typical structural characteristics found in various non-LTR families are internal pol II promoters and 3′ ends containing AATAAA polyadenylation signals, poly (A) tails, or simple tandem repeats. Target Primed Reverse Transcription has been proposed as the mechanism of retrotransposition for R2 of Bombyx mori, and this may be generally true for all non-LTR elements (Luan et al., 1993; Eickbush, 2002; Eickbush and Jamburuthugoda, 2008). Because they transpose by Target Primed Reverse Transcription, some non-LTR retrotransposons could rely rather heavily on host DNA repair mechanisms. This relationship with the host may give non-LTR retrotransposons some flexibility with regard to the domains required in an autonomous element (Eickbush and Malik, 2002). Some non-LTR retrotransposons, such as R2, are site-specific, because their endonucleases make precise cleavage at specific targets (Eickbush, 2002).
SINEs SINEs are generally between 100 and 500 bp long. Unlike LTR and non-LTR retrotransposons, SINEs do not have any coding potential. SINEs may have been borrowing the retrotransposition machinery from autonomous non-LTR retrotransposons, which may be facilitated by similar sequences or structures at the 3′ ends of a SINE and its "partner" non-LTR retrotransposon (Ohshima et al, 1996; Okada and Hamada, 1997; Dewannieux et al., 2003; Dewannieux and Heidmann, 2005). Unlike non-LTR retrotransposons that use internal Pol II promoters, SINE transcription is directed from their own Pol III promoters which are similar to those found in small RNA genes. SINEs can be further divided into three groups based on similarities of their 5′ sequences to different types of small RNA genes. Elements such as the primate Alu family share sequence similarities with 7SL RNA (Jurka, 1995), while most other SINEs belong to a different group that share sequence similarities to tRNA molecules (Adams et al., 1986; Okada, 1991; Tu, 1999; Tu et al, 2004; Luchetti and Mantovani, 2009; Xu et al, 2010). Recently, a new group of SINEs, named SINE3, have been discovered in the zebrafish genome, which share similarities to 5S rRNA (Kapitonov and Jurka, 2003a). Some non-LTR retrotransposons tend to generate truncated copies due to incomplete reverse transcription during cDNA synthesis. Although these short copies of RNA-mediated TEs are also called SINEs (Malik and Eickbush, 1998), they should not be confused with the true SINEs that use Pol III promoters.
Class II DNA-Mediated TEs
DNA-mediated TEs include cut-and-paste DNA transpo-sons (Figure 3), miniature inverted-repeat TEs (MITEs; e.g., Tu, 2001a; Coates et al., 2010a), Helitrons (Kapitonov and Jurka, 2001, 2003a; Thomas et al., 2010a), and a recently discovered group called Mavericks or Polintons (Feschotte and Pritham, 2005; Kapitonov and Jurka, 2006; Pritham et al, 2007). All Class II TEs transpose directly from DNA to DNA, and no RNA intermediate is involved (Craig, 2002).
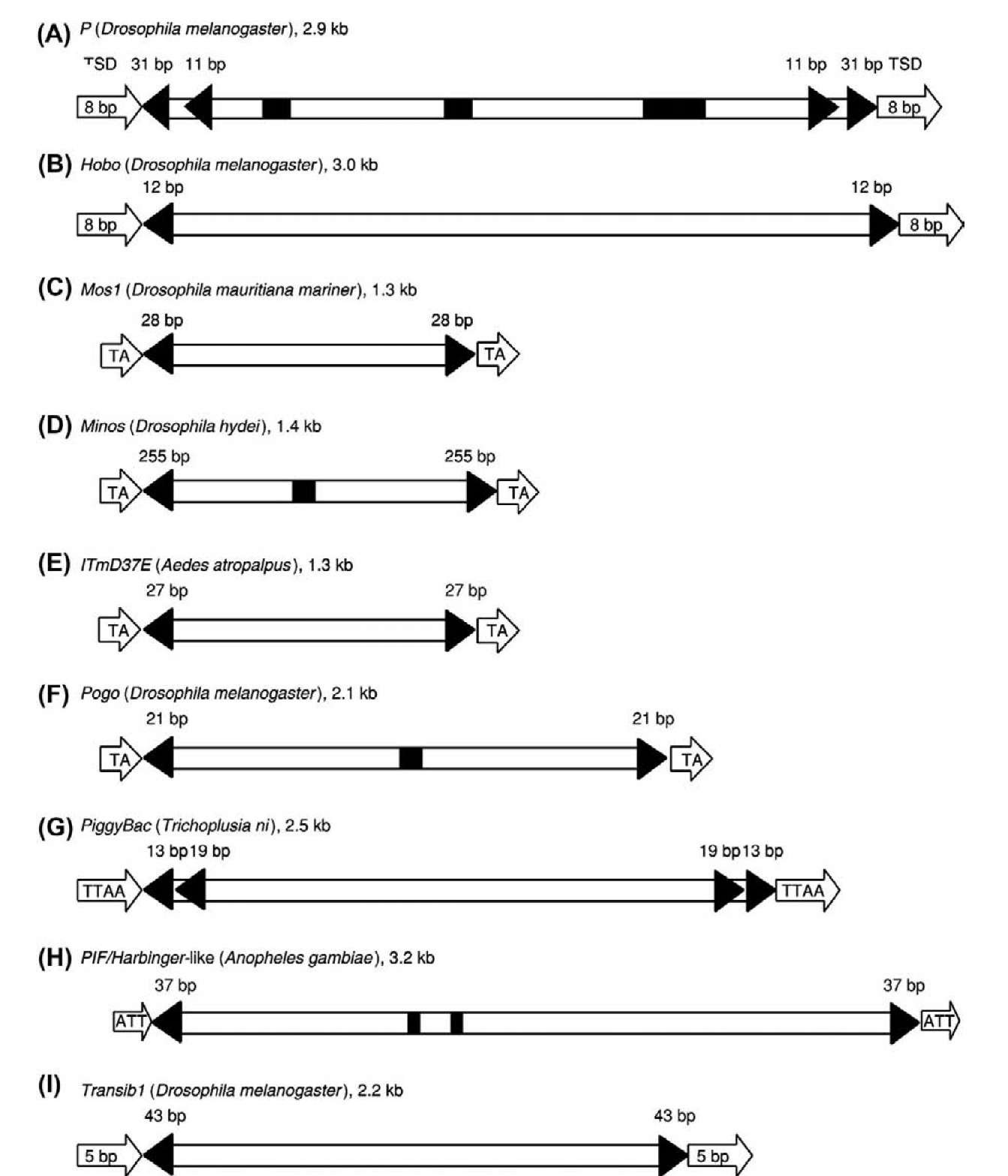
Cut-and-paste DNA transposons DNA transposons such as P, hobo, and mariner are usually characterized by 10- to 200-bp terminal inverted repeats (TIRs) flanking one or more open reading frames that encode a transposase. They usually transpose by a cut-and-paste mechanism, and their copy number can be increased through a repair mechanism (Finnegan, 1992; Craig, 2002; Zhou et al, 2004; Hickman et al, 2005; Mitra et al., 2008; Richardson et al., 2009). As shown in Figure 3, cut-and-paste DNA transposons can be subdivided into several families or superfamiles according to their transposase sequences. The families/superfamilies that have been found in insects include IS630-Tc1-mariner, hAT, Merlin, piggyBac, PIF/Harbinger, P, and Transib (Shao and Tu, 2001; Robertson, 2002; Kapitonov and Jurka, 2003a; Feschotte, 2004). These families/ superfamilies are also characterized by TSDs of specific sequence or length.
MITEs Miniature inverted-repeat TEs (MITEs) are widely distributed in plants, vertebrates, and invertebrates (Oosumi et al., 1995; Wessler et al., 1995; Smit and Riggs, 1996; Tu, 1997, 2001a; Yang et al., 2009; Coates et al., 2010b). Most MITEs share common structural characteristics, such as TIRs, small size, lack of coding potential, AT richness, and the potential to form stable secondary structures (Wessler et al., 1995). MITEs may have been "borrowing" the transposition machinery of autonomous DNA transposons by taking advantage of shared TIRs (MacRae and Clegg, 1992; Feschotte and Mouches, 2000a; Zhang et al., 2001). An alternative hypothesis suggests that they may transpose by a hairpin DNA intermediate produced from the folding back of single-stranded DNA during replication, which may better explain how MITEs could achieve immensely high copy numbers in some genomes (Izsvak et al., 1999). However, more recent evidence clearly favors cross-mobilization by autonomous transposons, and suggests that internal features of MITEs may help them achieve high transposition activity (Yang et al., 2009). One obvious source of MITEs is internal deleted autonomous DNA transposons (Feschotte and Mouches, 2000a). In this case, MITEs are basically non-autonomous deletion derivatives of DNA transposons. Recent studies show that the similarities between many MITEs and their putative autonomous partners are restricted to the TIRs (Feschotte et al., 2003). Although subsequent loss of autonomous partners in the genome remains a possible explanation for the lack of internal sequence similarity between MITEs and their putative autonomous partners, two other explanations are perhaps more plausible. First, MITEs could originate de novo from chance mutation or recombination events resulting in the association of TIRs flanking unrelated segments of DNA (MacRae and Clegg, 1992; Tu, 2000; Feschotte et al, 2003). Alternatively, these MITEs could originate from abortive gap-repair following the transposition of DNA transposons, which has been shown to occasionally introduce transposon-unrelated sequences (Rubin and Levy, 1997).
Helitrons: the rolling circle transposons Helitrons and related transposons have recently been discovered in insects and plants, which appear to use a rolling-circle mechanism of transposition (Le et al., 2000; Kapitonov and Jurka, 2001, 2003b; Coates et al., 2010b).
Figure 3 Structural characteristics of representative cut-and-paste DNA transposons in insects. The name of each representative transposon, its host species, and its approximate length are shown as the heading of each panel. Open arrows indicate target site duplications (TSDs); filled triangles indicate terminal and subterminal inverted repeats. The lengths of these inverted repeats are marked. Exons are shown as open boxes, and introns are shown as filled black boxes. 5′ and 3′ untranslated regions are not shown. The elements are not drawn to scale.
Instead of cut-and-paste transposase, Helitrons encode proteins similar to helicase, ssDNA-binding protein, and replication initiation protein. These proteins facilitate the rolling-circle replication of Helitrons, a mechanism previously described for the bacterial IS91 transposons (Garcillan-Barcia et al., 2002).
Polintons/Mavericks: giant DNA transposons that encode integrase as well as DNA polymerase A group of large DNA transponsons that were first discovered in Tetrahymena (Wuitschick et al., 2002) have recently been shown to be broadly distributed in metazoan, fungi, and various single-cell eukaryotes (Feschotte and Pritham, 2005; Kapitonov and Jurka, 2006; Pritham et al., 2007). These elements are either called Politons or Mavericks, and share several features, including 6-bp TSD, long TIRs, and coding sequences for integrase, DNA polymerase, and a few other proteins. They are sometimes 20 kb in length, and appear to be related to adenoviruses, bacteriophages, and eukaryotic linear plasmids. It is proposed that an excised Polinton/Maverick can self-replicate with its own polymerase and integrate into the genome using its integrase.
Related Topics
Foldback elements Drosophila Foldback elements are characterized by very long inverted repeats (Truett et al., 1981). It is not known how Foldback elements transpose, although the presence of long inverted repeats indicates a possible DNA-mediated mechanism. Some researchers group Foldback elements as a distinct class, namely Class III (Kaminker et al., 2002).
What is a family? Before moving ahead with discussions on the discovery and diversity of insect TEs in the next two sections, it may be helpful to clarify the use of the term "family" in the context of TEs. The term "family" is often used to refer to a group of related TEs in diverse organisms that usually share conserved amino acid sequences in their transposase or reverse transcriptase. The mariner family is such an example. A TE also consists of many copies generated by transposition events in a genome. Therefore, these related copies are sometimes also referred to as a family. Some families consist of multiple distinct groups that are subdivided into subfamilies. Obviously, relatedness is a relative concept in evolution. A working definition is needed in each case until a universal family/subfamily definition is developed.