Mechanism of Action of miRNAs
The functional role of a miRNA is ultimately characterized by its effects on the expression of target genes. Currently, the regulatory mechanisms involving miRNAs are related to mRNA cleavage or translational repression by binding to complementary sites usually located on the 3′ UTR region of the mRNA (Carrington and Ambros, 2003; Lai, 2003; Ambros, 2004; Bartel, 2004). In contrast to the inhibitory effects, miRNAs can also stimulate the expression of target genes by upregulation of translation (Vasudevan et al., 2007; Orom et al., 2008). Moreover, miRNAs can also control cell fate by binding to heterogeneous ribonu-cleoproteins and lifting the translational repression of their target mRNAs; in this way, miRNAs act through a sort of decoy activity that interferes with the function of regulatory proteins (Beitzinger and Meister, 2010; Eiring et al., 2010). The present section, however, will emphasize the more widespread mechanisms, leading to mRNA translational repression, which start when the miRNA binds to Ago-1 protein and with the assembly of the RISC (Figure 2).
Argonaute Loading
The Argonaute (Ago) family can be divided into two subfamilies: the Piwi subfamily and the Ago subfamily. Piwi proteins are involved in transposon silencing, and are especially abundant in germ-line cells. Ago-subfamily proteins play key roles in post-transcriptional gene regulation by interacting with siRNAs (see above) and miRNAs, as detailed below.
After Dicer-1-mediated cleavage, the miRNA duplex binds to an Ago-1 protein in the RISC. To form an active RISC, the miRNA duplex has to unwind because only the mature miRNA binds to the Ago-1 protein, whereas the miRNA* is released. In human cells, miRNAs with a high degree of base-pairing in their pre-miRNA hairpin stem are initially processed by Ago-2, which cleaves the 3′ arm of the hairpin (that is, the miRNA* strand) in the middle, thus generating a nicked hairpin (Diederichs and Haber, 2007). In this case, Ago-2 acts before Dicer-1-mediated cleavage and facilitates miRNA duplex dissociation, the removal of nicked strand, and the activation of RISC. These findings elucidated the crucial role of Ago proteins not only during RISC formation, but also in relation to the mechanism that determines which of the two strands will become the survivor mature miRNA.
Identification of the target mRNA by the RISC is based on the complementarity between the mature miRNA and the target mRNA site, and the degree of complementarity determines whether the target mRNA is degraded, destabilized, or translationally inhibited. Binding to Ago-1 greatly enhances miRNA stability, and although little is known about the half-life of individual miRNAs, it is clear that Ago-1 is a limiting factor for endogenous miRNA accumulation due to its protective function.
In D. melanogaster, miRNA* strands may accumulate bound to Ago-2, a protein initially thought to act exclusively in the siRNA pathway. Whether miRNA* binds to Ago-1 or to Ago-2 depends on the miRNA duplex structure, ther-modynamic stability, and the identity of first 5′-end nucleotide — i.e., miRNA sequences beginning with cystidine will bind to Ago-2, whereas those beginning with uridine will bind to Ago-1 (Ghildiyal et al., 2010). A number of observations indicate that some miRNA* plays a role in the regulation of gene expression. These observations include that: (1) miRNA* 5′ ends are more defined than their 3′ ends, thus suggesting that there is a seed region involved in regulatory functions (Ruby et al., 2007b; Okamura et al., 2008; Seitz et al., 2008); (2) many miRNA* sequences are evolutionarily conserved (Okamura et al., 2008); (3) in D. melanogaster (Ruby et al., 2007b) and in the basal insect B. germanica (Cristino et al., 2011), tissue concentration of some miRNA* is higher than that of the corresponding miRNA partner.
Repression of Protein Translation
The RISC is the key element that regulates gene expression by repressing protein translation. The first step after RISC formation is the recognition of the target mRNA, mainly through the seed sequence. A number of studies have demonstrated the importance not only of the seed, but also of the whole 5′ region of the miRNA during the interaction with the target mRNA. According to Brennecke and colleagues (2005), there are two categories of miRNA target sites in mRNAs. The first is called the "5′ dominant site," and occurs when there is a near perfect base-pairing in the 5′ end of the miRNA; this category can be subdivided into "canonical" (when both 5′ and 3′ ends have strong base-pairing with the miRNA site) and "seed" (when only the 5′ region presents consistent base-pairing). The second category is called "3′ compensatory," and occurs when base-pairing between the miRNA seed sequence and its corresponding sequence in the target mRNA is weak, and thus a stronger base-pairing in the 3′ region exerts a sort of "compensating" effect.
Initial experiments in C. elegans showed that the mi RNAs lin-4 and let-7 repress their respective target mRNAs through interactions with miRNA sites in the 3′ UTR. Subsequently, many other cases of miRNA binding sites in the 3′ UTR of mRNAs were reported, leading to the presumption that this was a general rule. However, recent findings have revealed that miRNAs can repress mRNAs through sites located in the open reading frame (ORF) or in the 5′ UTR (Lee et al., 2009).
The action of RISC on target mRNAs may proceed through different mechanisms. One of them involves post-initiation repression, as shown by experiments carried out in C. elegans where lin-4 inhibits the translation of lin-14 mRNA without reducing the mRNA levels and without affecting the shifting of polysomes, thus suggesting that the inhibition of mRNA translation occurs at the elongation step (Wightman et al, 1993; Olsen and Ambros, 1999; Lee et al., 2003). Other details accounting for this mechanism of action have been reported, and a model has been proposed describing the inhibition of ribosome elongation, the induction of ribosome drop-off, and the facilitation of nascent polypeptides proteolysis (Fabian et al., 2009).
The second mechanism of RISC action is the acceleration of target mRNA destabilization, involving: (1) decap-ping of the m(7)G cap structure in the 5′ end; and/or (2) deadenylation of poly A tail during the initial step of translation (Humphreys et al., 2005). A number of reports using different experimental models have supported this second mechanism; for example, in zebrafish embryos and mammalian cells, miRNAs in the RISC accelerate mRNA deadenylation, which leads to fast mRNA decay (Figure 5) (Giraldez et al., 2006; Wu et al, 2006). In Drosophila cells both deadenylation and decapping require GW182 protein, CCR4 : NOT deadenylase, and the DCP1 : DCP2 decapping complexes. Depletion of GW182 in Drosophila cells leads to alteration of mRNA expression levels. However, in Ago-1depleted cells, GW182 can still silence the expression of target mRNAs, thus indicating that GW182 acts downstream of Ago-1, and that it is a key component of the miRNA pathway (Behm-Ansmant et al., 2006a).
Processing Bodies and mRNA Storage
In many cases, the last step of miRNA action involves the processing bodies (P-bodies), which are discrete cyto-plasmic aggregates that contain enzymes associated to mRNA decay, such as CCR4 : NOT complex (deadenylase), DCP1 : DCP2 complex (decapping), RCK/p54, and eIF4ET (general translational repressors). The aforementioned GW182 is additionally required for P-body integrity. Apparently, P-bodies are the place where RISC delivers its target mRNA to be degraded or to be stored (Figure 5). In human cells, for example, miR-122-repressed mRNAs that are maintained in P-bodies can be released from them under stress conditions, and subsequently be recruited by polysomes (Bhattacharyya et al., 2006). Behm-Ansmant and colleagues (2006b) have proposed a model where RISC binds to target mRNA through interactions with miRNA and Ago-1, and recruits GW182, which labels the transcript as a target for decay via deadenylation and decap-ping.
Figure 5 Once the miRISC is formed, the target mRNA can be taken to a special region of cytoplasm known as the P-body, where it will be degraded after decapping and deadenylation, or maintained in the P-body until released from it and recruited to polysomes.
Ago-1 and Ago-2 proteins have also been detected in P-bodies (Liu et al., 2005), thus suggesting that both siRNA and miRNA pathways may end in these structures. Nevertheless, this does not mean that P-bodies are crucial for the functioning of these pathways, given that disruption of P-bodies after depletion of Lsm1, which is a key component of them, elicits a dispersion of Ago proteins into the cytoplasm, but does not affect siRNA and miRNA pathways (Chu and Rana, 2006).
Identification of miRNAs in Insects
Since the discovery of lin-4 and let-7 in the nematode C. elegans, a remarkable diversity of miRNAs has been reported in the genomes of various organisms, including insects, plants, viruses, and vertebrates (http:// www.mirbase.org). In insects, research on miRNAs was initially limited to D. melanogaster, but the availability of sequenced genomes from different species, as well as the development of new bioinformatic tools, has allowed the performance of systematic predictions of miRNAs in silico. Accordingly, computational methods based on the evolutionary conservation of genomic sequences and their ability to fold into stable hairpin structures have been applied to species with sequenced genomes, such as a number of nematodes, arthropods, and vertebrates (Table 1). Moreover, the development of novel techniques for directional cloning of small RNAs has led to the identification of many other miRNAs (Lagos-Quintana et al., 2001; Lau et al., 2001; Lee and Ambros, 2001).
Nevertheless, the greatest progress came with the advent of high-throughput sequencing technologies and computational methods. Those technologies confirmed most of the miRNA predicted in silico in species with the genome reported, made it possible to find new and unexpected miRNAs, and contributed to the discovery de novo of miRNAs in species without the genome sequenced. Therefore, a consistent catalog of miRNAs is now available not only in drosophilids, but also in a selection of species, such as the malaria mosquito (A. gambiae), the yellow fever mosquito (Aedes aegypti), the pea aphid (A. pisum), the vector of West Nile virus (Culex quin-quefasciatus), the jewel wasp (Nasonia vitripennis), the migratory locust (Locusta migratoria), the honey bee (Apis mellifera), the flour beetle (T. castaneum), the silkworm (Bombyx mori), and the German cockroach (B. germanica) (http://www.mirbase.org; http://www.ncbi.nlm.nih.gov/geo) (Griffiths-Jones, 2006). Both approaches, based on computational methods and high-throughput sequencing, are discussed below.
Computational Methods
The most efficient computational methods for finding miRNA candidates were described in C. elegans (MiRscan) (Lim et al., 2003a) and D. melanogaster (miRseeker) (Lai et al., 2003). Both methods share conceptual similarities, such as structural and sequence similarity. MiRscan produces an initial set of candidates by sliding a 110-nucleotide window across the C. elegans genome and folding those segments that are filtered by the free energy and duplex length. Homologous hairpins are then identified by WU-BLAST in an additional genome which creates a reference set defining the standard features that will finally be used to score and rank all candidate hairpins. Nevertheless, MiRscan was not able to identify more than 50% of the previously known C. elegans miRNAs (Lim et al., 2003a). miRseeker was found to be more efficient at identifying genuine miRNAs in two fly species (D. melanogaster and Drosophila pseudoobscura) by taking into account the conservation across the hairpin (Lai et al., 2003). The method begins by identifying orthologous intergenic and intronic regions of those two fly genomes, and then folding those conserved sequences to identify and score the hairpin structures.
Table 1 Algorithms Developed for miRNA Identification
|
Program |
Strategy |
Species group |
Authors/year |
|
Grad et al. |
RB |
Nematodes |
Grad et al., 2003 |
|
MiRScan |
RB |
Nematodes, vertebrates |
Lim et al., 2003a, 2003b |
|
miRseeker |
RB |
Insects (flies) |
Lai et al., 2003 |
|
Berezikov et al. |
RB |
Human |
Berezikov et al., 2005 |
|
miPred |
RB |
Human |
Jiang et al., 2007 |
|
miRAlign |
RB |
Metazoan |
Wang et al., 2005 |
|
ProMIR |
HMM |
Human |
Nam et al., 2005 |
|
BayesMiRNAFind |
NB |
Nematodes, mammals |
Yousef et al., 2006 |
|
One-ClassMirnaFind |
SVM,NB |
Human, virus |
Yousef et al., 2008 |
|
mirCoS-A |
SVM |
Mammals |
Sheng et al., 2007 |
|
mir-abela |
SVM |
Mammals |
Sewer et al., 2005 |
|
triplet-SVM |
SVM |
Human |
Xue et al., 2005 |
|
RNAmicro |
SVM |
Metazoan |
Hertel and Stadler, 2006 |
|
miPred |
SVM |
Human |
Ng and Mishra, 2007 |
|
MiRFinder |
SVM |
Human, virus |
Huang et al., 2007 |
The criteria for hairpin evaluation derive from a reference set of known miRNA genes of the two Drosophila species. The length of the hairpins and their minimum free energy were first evaluated, and then the distribution of divergent nucleotides was considered to score the candidates. The metrics consist in penalizing divergences depending on where they occur in the pre-miRNA hairpin, as the miRNA arm would tolerate less mutations than the miRNA* arm, which, by itself, would not tolerate more mutations than those observed in the loop region (Lai et al, 2003).
The establishment of guidelines for the experimental validation and annotation of novel miRNA candidates became obviously necessary with the increasing quantity of miRNA genes being identified in various species (Ambros et al., 2003). Thus, an initiative for organizing the information available on miRNA genes was then developed, leading to a database (miRBase, http://www.mirbase.org) where all data regarding miRNA sequences, targets, and gene nomenclature are deposited (Griffiths-Jones et al., 2008).
The large amount of miRNA data available in databases led to the development of a second generation of algorithms based on machine-learning methods. The approach consists in a learning process that identifies the most relevant characteristics and rules from a positive set of miRNA hairpins. Various machine-learning algorithms have been used for miRNA discovery (Table 1), the most common being Naive Bayes (Yousef et al., 2006), support vector machines (Yousef et al., 2008 and references therein), hidden Markov models (HMM) (Nam et al., 2005), genetic programming (Brameier and Wiuf, 2007), and random walks (Jiang et al., 2007).
All these methods contributed somehow to the identification of new miRNAs, despite considerable differences in their trade-off between specificity and sensitivity. The criteria used in all of them were based on actual knowledge of the miRNA biogenesis, and features identified from known miRNAs conserved in at least two species. Indeed, there must be a great number of non-conserved miRNA genes still to be discovered, which may have characteristics and expression profiles substantially different from those of canonical miRNAs. However, the development of a new generation of sequencing technologies is changing the way of thinking about scientific approaches in all fields of biological sciences (Metzker, 2010), including the strategies to find new miRNAs in any species, even those whose genome is not sequenced yet.
High-Throughput Sequencing
Deep-sequencing technologies have created a new paradigm in detecting low-expression or tissue-specific miRNAs, as well as non-canonical and species-specific ones. The most effective algorithms published so far are miRDeep (Yang et al., 2010), MIReNA (Mathelier and Carbone, 2010), and deepBase (Friedlander et al., 2008). Despite varying slightly in their workflow, their general strategy is similar, combining mapping and filtering sequences based on genome annotation, sequence and structure patterns, and properties of miRNA biogenesis.
The identification of miRNAs through deep-sequencing methods is rapidly increasing the catalogs of small RNA sequences for many species from a variety of taxonomic groups. Currently, all deep-sequencing datasets are deposited in the GEO (Gene Expression Omnibus) database at the NCBI (National Center for Biotechnology Information; http://www.ncbi.nlm.nih.gov/geo). At the date of writing (January 2011), there are at least 193 studies of high-throughput sequencing of small RNAs from different eukaryotic species in the GEO database. Table 2 shows the 14 insect species in the GEO database, and the number of records for each.
Most of the 14 insect species included in Table 2 have the genome sequenced, or at least have a closely related species with an available genome (e.g., A. albopictus and C. quinquefasciatus). Two species, L. migratoria and B. germanica, have no genome sequence available, and identification of miRNAs from deep-sequencing data becomes challenging because none of the methods mentioned above were designed to analyze deep-sequencing data without using a genome sequence as a reference, and the diversity of small RNA types is remarkably high.
Table 2 Insect Species and Number of Records Found in the GEO Database Related to Studies of miRNA Identification
|
Order |
Species |
Number of records |
|
Diptera |
||
|
Drosophila melanogaster Drosophila simulans Drosophila erecta Drosophila pseudoobscura Drosophila virilis Aedes albopictus Culex quinquefasciatus |
21 1 1 1 1 1 1 |
|
|
Lepidoptera |
||
|
Bombyx mori |
2 |
|
|
Hymenoptera |
||
|
Camponotus floridanus Harpegnathos saltator Apis mellifera |
1 1 1 |
|
|
Hemiptera |
||
|
Acyrthosiphon pisum |
1 |
|
|
Orthoptera |
||
|
Locusta migratoria |
1 |
|
|
Dyctioptera |
||
|
Blattella germanica |
1 |
|
However, strategies that can identify previously described miRNAs, as well as novel miRNAs on the basis of the number of reads and hairpin features, have recently been proposed (Wei et al., 2009). Genome-independent approaches for miRNA discovery show that we still have a poor understanding of the small RNA world and its regulatory mechanisms in the cell. For example, in the locust L. migratoria (Wei et al., 2009) and in the cockroach B. germanica (Cristino et al., 2011), sequence read numbers corresponding to miRNA*s were higher than those corresponding to the mature miRNA. Another original finding has been reported in Drosophila species (Berezikov et al., 2010), where some miRNA precursors seem not to be processed by RNase III only, given that the usual one- to two-nucleotide 3′ overhang does not occur in some sequences represented by a high number of reads.
miRNA Classification
As stated above, identification efforts have led to the description of an impressive number of miRNAs in animals, plants, green algae, fungi, and virus (Griffiths-Jones et al., 2008), and different attempts to classify such a high diversity into families based on structural coincidences have been carried out. As the pattern of nucleotide substitution in miRNA genes is apparently shaped by selective pressures, and considering that the seed is the most important region from a functional point of view (Brennecke et al, 2005; Bartel, 2009), miRNA classification is based on this region. Regarding metazoans, 858 miRNA families are deposited in the miRBase database (v16.0) (Griffiths-Jones et al., 2008), and 254 (30%) of these families are found in at least five species. These records will change with further high-throughput sequencing experiments, but present data indicate that most of the miRNA families (a total of562) are found in vertebrates, followed by insects (178 families reported), and then by other metazoan that are phyllogenetically more basal, such as cnidaria, porifera, hemichordata, echinodermata, urochordata, cephalochor-data, and nematoda (118 families in all).
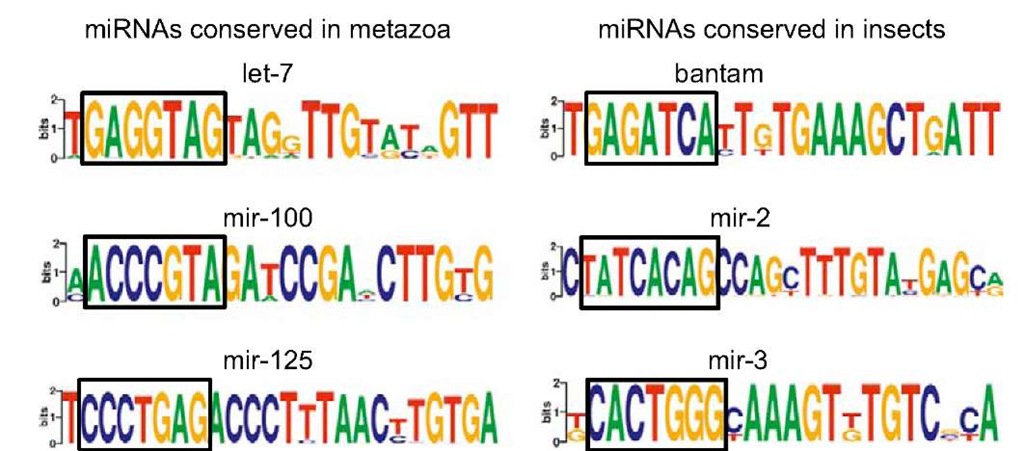
The seed region can be more or less conserved in different miRNA families. A good example of a well-conserved seed region is observed in the miRNAs miR-100, miR-125, and let-7 (Behura, 2007). As stated above (see also Figure 3), these three miRNAs are often coded by the same polycistronic pri-miRNA that has a conserved organization from invertebrates to vertebrates, which suggest that it is an ancestral pri-miRNA. As expected, the seed region of these miRNAs is highly conserved (Figure 6). There are insect-specific miRNA families whose seed region is also very well conserved, as for instance bantam miR-2 and miR-3 (Figure 6). The conservation of the seed region occurs not only among paralogous sequences, resulting from intraspecific gene duplication, but also among orthologous sequences arising from speciation events. Of note, the conservation of the seed region is critical for the recognition of mRNA targets, thus the classification of miRNAs into families on the basis of the seed not only contains structural information, but may also reflect functional regularities.
Target Prediction
In animals, the functional duplexes miRNA : mRNA can occur in a variety of structures where short complementary sequences can be interrupted by gaps and mismatches (Brennecke et al., 2005; Bartel, 2009). Thus, most computational methods have been developed to find target sequences based on the complementarity between the miRNA seed sequence and the mRNA sequence. Several computational approaches estimate the likelihood of miRNA : mRNA duplex formation, mainly based on sequence complementarity, thermodynamic stability, and evolutionary conservation of the sequence among species (Table 3).
Figure 6 Conservation of miRNA genes on the region corresponding to mature miRNAs in metazoan and insects. The sequence logo is constructed based on the alignment of various miRNA sequences representing the level of nucleotide conservation in each position. The squares indicate the canonical seed regions located at nucleotides 2-8.
Machine learning approaches are also used for miRNA target identification. These methods usually combine one or more of the traditional procedures (seed complementarity, thermodynamic stability, and cross-species conservation) with more elaborated probabilistic models (Table 3). Also, a new generation of algorithms is integrating high-throughput expression data and computational predictions (Huang et al., 2007; Hammell et al., 2008; van Dongen et al., 2008; Wang and El Naqa, 2008; Bandyopadhyay and Mitra, 2009; H. Liu et al., 2010; Sturm et al, 2010).
To date, miRNA target prediction has been mainly performed by computational approaches, and large numbers of targets have been predicted for most species with the genome sequenced (Bartel, 2009). As a general figure, predictions have suggested that a single miRNA can target 200 mRNAs on average in vertebrates (Krek et al., 2005), whereas in D. melanogaster a single miRNA may regulate 54 genes on average (Grun et al., 2005).