Proteomics
Proteomics is the study of all proteins present in an organism, and deals with their quantification, identification, and modifications that alter their function. While statistically significant changes in mRNA levels are usually correlated with changes in protein levels, individual proteins can change drastically with little significant correlation at the mRNA level (Bonaldi et al., 2008). Cellular protein abundance is controlled through many different mechanisms. These mechanisms include translational efficiency based in part on regulatory sequences in the 5′ and 3′ untranslated regions of mRNA, and protein degradation through ubiquitination and the 28S proteasome pathway. Post-translational modifications and the presence of interacting partners often alter the function or the functional capacity of a protein.
Modern proteomics relies heavily on mass spectro-metry (MS). Mass spectrometry devices measure the mass-to-charge ratio of peptide ions. Mass spectrom-etry can be used for protein quantitation, identification, and sequencing, and determining the presence of post-translational modifications. Two broad MS strategies, the bottom-up approach and the top-down approach, vary on whether proteolytically digested peptides are analyzed, or the entire protein is sequenced. In the bottom-up approach, peptides of interest are often separated on a two-dimensional (2D) gel, extracted, digested into smaller fragments via trypsin proteolysis, and analyzed by MS. Often, the amino acid sequence and corresponding mass (M) to charge (z) (or M/z) ratio between two tryp-sin cut sites are sufficient to identify a protein. The mass of the digested peptide is compared against a sequence database containing all genomic open reading frames and their calculated masses. This approach is also known as peptide mass fingerprinting. In the top-down approach, a whole protein can be sequenced using tandem MS, or MS/MS. Tandem MS measures the M/z ratio of a protein ion before fragmentation, and the resulting amino acid or peptide ions after fragmentation. Finally, in shotgun proteomics, a large number of proteins are first digested, then separated by HPLC, and finally analyzed, often by tandem MS.
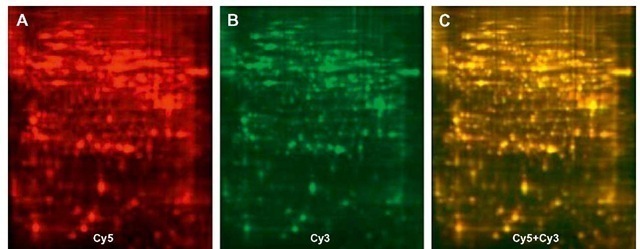
Proteins need to be separated before MS analysis, and separation is usually accomplished by Liquid Chromatog-raphy (LC), High Performance LC (HPLC), or 2D gel electrophoresis. In order to identify proteins with varying abundance between two treatment groups, differential gel electrophoresis (DIGE) can be used, and DIGE can be followed by Matrix Assisted Laser Desorption-Time of Flight (MALDI-TOF) MS analysis (MALDI, matrix-assisted laser desorption/ionization, or TOF, time-of-flight mass spectrometer). In DIGE, proteins from two treatment groups are extracted, mixed with different colored dyes, usually CY3 and CY5, and subsequently run on a 2D polyacrylamide gel which separates proteins based on size and isoelectric focusing point (Gorg et al., 2004) (Figure 5). Changes in protein expression can be inferred from changes in the color and intensity of "spots" on the gel, which usually represent one protein. Because the CY3 emission spectrum is in the green range and CY5 fluoresces in the red spectrum, proteins that are equally present in both treatments appear as yellow spots, while those that are up- or downregulated appear as orange spots, and those present in only one treatment group appear red or green. Algorithms have been developed to quantify the spot intensity and protein quantity (Gorg et al., 2000; Herbert et al., 2001; Patton and Beechem, 2002), but the identity of the protein remains unknown and the spots must therefore be subjected to MS. Similar to mRNA expression measurement, changes in protein levels between two treatment groups must be analyzed statistically for significance.
Differential gel electrophoresis may be followed by pep-tide mass fingerprinting, or PMF. MALDI-TOF is often coupled to trypsin proteolysis, a bottom-up approach, which is simpler and has greater throughput than MS/ MS. After extracting a spot from a 2D gel, the protein must be digested with trypsin, ionized, and finally introduced into the MS device. Introduction can be accomplished by MALDI, or electrospray ionization, and M/z detection may be accomplished by a Time of Flight (TOF) detector. After digestion, the peptide spot is added to a protective matrix. Next, a laser beam converts the protein from a solid molecule into a gas-phase ion with minimal damage to the protein. The matrix protects the protein by absorbing most of the laser energy, and ionizes the protein through a poorly understood mechanism which may involve charge transfer (Knochenmuss, 2006). Mixtures of proteins or digested peptides are further separated by the action of the laser, which only ionizes portions of the matrix, thus reducing the chance of different fragments entering the TOF analyzer at once.
In a typical MALDI-TOF analysis, the laser-based ionization of a peptide fragment accelerates ions into a vacuum where an electrical field is applied perpendicular to the direction of ionization. In this way, all ions have the same potential energy and velocity of zero in the axis towards the mass detector. Potential energy in the form of voltage is equally applied to the ions, which causes them to accelerate towards the TOF detector. Since the voltage applied is uniform, the velocity at which the ions travel is dependent on their mass and charge. The distance traveled from the field to the detector is constant for the same MS instrument. Time is experimentally measured between application of the electric field and arrival at the mass detector. Time is therefore proportional to mass and charge.
Figure 5 Two-dimensional differential in-gel electrophoresis (2D-DIGE) images of insecticide-susceptible (Cy5-labeled, Panel A) and resistant (Cy3-labeled, Panel B) SF-21 cells treated with insecticide. Panel C is an overlay of the two images. Equal amounts of protein in both cell lines appear yellow (C) and the proteins present in only resistant cells appear green (B), while only susceptible cells appear red (A).
The resulting data can often be used to identify proteins. However, the amino acid sequence cannot be determined, since the final peptide masses could result from a number of amino acid combinations. For PMF, a genomic sequence database is required to match the digested peptide mass against known proteins and open reading frames. Tandem mass spectrometry is a popular application for the identification, quantitation, and de novo sequencing of proteins. Protein mixtures need not be previously digested enzymatically, and some separation can be achieved by a preliminary mass analyzer inside the MS device. One type of mass analyzer is a quadropole ion trap, which uses DC and AC electrical fields and RF frequencies to trap or capture entering peptide ions. By changing the AC field frequency, pep-tides of different M/z ratios can be selected, and this is therefore the first M/z analysis, or MS in tandem MS, or MS/MS. In a typical peptide-sequencing experiment, an isolated, selected protein may be fragmented into smaller peptides or even amino acids. Fragmentation may be accomplished by collision-induced dissociation (CID), where the protein is bombarded with neutral ions. Fragmentation can occur at three predictable spots on the protein backbone. The smaller peptides are then caught in a final mass analyzer before detection. The final mass analyzer may be a TOF analyzer or a more sophisticated analyzer. Proteins for tandem MS can be enriched for post-translational modifications, or separated through a number of chromatographic steps. HPLC is often used to separate proteins immediately upstream of MS/MS, and when LC separation is performed on an entire pro-teome the technique is called shotgun proteomics. Ion-ization and introduction into MS/MS analyzers from LC separation can often be achieved by electrospray ioniza-tion, where the LC solvent evaporates and causes ioniza-tion without fragmentation.
Sample Protein Labeling and Separation
Quantification of protein expression changes between two unlabeled treatments is not possible using shotgun pro-teomics, because the identical proteins have identical M/z ratios. A number of techniques have been developed to uniquely label proteins from a treatment without altering their function. Most of these techniques are applicable to cell culture, while one has been applied to two whole organisms. Stable isotopic labeling in cell lines is a labeling technique that allows protein quantitation between two treatments (Mann, 2006). Cell cultures are supplemented with either natural amino acids (light chain) or stable isotope labeled amino acids which are then incorporated into proteins (Ong et al., 2002). Deuterium/hydrogen, 12C/13C, and 14N/15N are commonly used non-radioactive isotopes that can be combined to accommodate greater sample numbers. MS is sensitive enough to detect the small mass changes.
Other quantification methods have been developed that label the protein after extraction from the cell. Isotope Coded Affinity Tag (ICAT) makes use of a label that reacts with cysteines, separated by a linker group that contains either deuterium (heavy) or hydrogen (light), and a biotin affinity tag. Proteins are extracted and enzy-matically digested, and cysteine containing peptides are purified using streptavadin, and finally subjected to MS (Gygi et al., 1999). Bonaldi et al. (2008) used SILAC (stable isotope labeling by amino acids in cell culture) to analyze the Drosophila S2 cell line proteome with the use of RNAi, and found that label incorporation did not affect protein expression. Interestingly, overall protein levels changed with little correlation to mRNA changes; however, when statistically significant changes occurred between knockdown and control, the mRNA change was highly correlated with changes in protein concentration. Only two animals have been successfully labeled using SILAC: the mouse and the fruit fly (Gygi et al., 1999; Sury et al., 2010).
Enrichment for PTM
Analyzing an entire proteome from two SILAC treatments and detecting post-translational modifications can be complex due to database searching with an increased number of mass ranges that could uniquely identify a protein. Enrichment may reduce complexity by focusing efforts on a smaller subset of interesting proteins. Antibody-based enrichment is one such method, and antibodies for PTMs can be purchased or custom-designed for specific needs. Examples of PTM antibodies include anti-phosphotyrosine/serine, anti-ubiquitin, etc. Samples may be digested with trypsin before enrichment to further decrease complexity and non-specific interactions. Antibody enrichment can be achieved by either immuno-affinity purification or immunoprecipitation (Zhao and Jensen, 2009). Efficacy may vary between these methods, based on the PTM of interest. Phosphotyrosine proteins fare better when immunoprecipitation is used (Schumacher et al. , 2007).
Applications of Proteomics
In parallel to genomics, proteomics provides a global view of protein profiles in an organism. Moreover, newly developed proteomics technologies allow for the deciphering of complicated biological systems, including cellular protein-protein interaction networks and various post-translational modifications. Proteomics technologies have been applied to study protein expression patterns among different insect developmental stages (Zhao et al., 2006; Li et al, 2007; Zhang et al., 2007; Chan and Foster, 2008; Li et al., 2009; Wu et al., 2009) and various insect tissues, such as reproductive tissues (Kelleher et al., 2009; Take-mori and Yamamoto, 2009), the nervous system salivary and silk glands (Zhang et al., 2006; Almeras et al., 2009), the cuticle (Holm and Sander, 1997), and hemolymph (Li et al., 2006; Furusawa et al., 2008a). Proteomics has been used to identify novel venom proteins (de Graaf et al., 2010) and salivary gland proteins (Oleaga et al., 2007; Carolan et al., 2009), as well as royal jelly proteins from the honey bee (Furusawa et al., 2008b; Li et al., 2008b; Yu et al., 2010). In addition, proteomics has been applied in studies on insect-plant and host-parasite interactions (Chen et al., 2005; Biron et al., 2005, 2006; Francis et al., 2006; An Nguyen et al., 2007). Interestingly, proteomic-based de novo gene discovery has been applied for identifying novel genes that are not predicted by genome annotation (Findlay et al., 2009). The development of powerful phosphoproteomics techniques enables large-scale identification of post-translational modifications, such as phosphorylation (Fu et al., 2009; Rewitz et al., 2009). Insecticide resistance (e.g., Cry toxins produced by the soil bacterium Bacillus thuringiensis) has become a serious problem that threatens Bt-based pest control and management. It is important to understand the mode of action of Cry toxins, especially the interaction between Cry toxins and host defense systems. Several studies have applied proteomics technologies to discover Cry binding proteins (McNall and Adang, 2003; Krishnamoorthy et al., 2007; Bayyareddy et al., 2009; Chen et al., 2009) and alterations of larval gut proteins between susceptible and resistant Indian meal moths (Candas et al., 2003).
Structural Genomics
Structural genomics is the study of the three-dimensional structure of all proteins from a particular organism through a combination of experimental determination and in silico modeling. The goal of structural genomics is set by some (Vitkup et al., 2001) as the ability to model 90% of the proteins within a genome through computational techniques using a much smaller number of carefully selected proteins representative of different protein families. Vit-kup’s survey concluded that, given the structural coverage in the Protein Data Bank (PDB, www.pdb.org; Berman et al. , 2002), only about 10% of the amino acids in a genome can be modeled. Based on the rate of 50 structures solved per week (Weissig and Bourne, 1999), and the observation that only 10 of these are non-redundant based on accepted definitions of protein families (Holm and Sander, 1997; Brenner and Levitt, 2000), a realistic application of structural genomics may lie decades in the future.
However, homology modeling is an effective tool for analyzing protein function, especially in the field of entomology. Insects represent a genetically diverse class of organisms, yet comparatively few insect protein structures have been solved to date. The time required to create an accurate homology model can be less than a week -sometimes even a day – and no specialized equipment is required. Models can yield information on ligand and substrate binding, their binding specificity, the evolutionary conservation of residues, the consequences of mutations in regard to pesticide resistance, and potential protein interactions, as well as elucidate targets of interest for further "wet" experiments.
Many of the limitations of modeling correlate with the template-target sequence identity and the subsequent difficulties in obtaining a correct alignment. For example, a protein with 70% sequence identity, or 70 amino acids the same in 100, may yield a target structure that is accurate enough for reasonable positioning of hydrogen atoms given a high-resolution template. Sequences with sequence identity as low as 20% could still be considered useful for many applications, especially when combined with comparative homology data. Docking ligands, pesticides, or drugs into these models is one such task.
Homology modeling has been applied to determine the substrate specificity for two different p450s in Anopheles gambiae which shared only 20% identity with their human template (Chiu et al., 2008). P450s are a class of proteins which chemically alter a wide range of substrates, including pesticides, through hydroxylation to facilitate excretion. Mutations in a voltage-gated sodium channel from the house fly Musca domestica have been mapped onto homologous structures from mammals, and used to elucidate the role of these mutations in pesticide resistance.The aryl hydrocarbon receptor, a bHLH PAS transcription factor which controls the expression of proteins related to carcinogen decay, was successfully modeled, and a conserved ligand-binding domain was found. Through structure-based mutagenesis, residues involved in binding the carcinogenic xenobiotic TCDD were successfully elucidated (Pandini et al., 2009). More generally, conservation of residues across evolu-tionarily diverse organisms, or between highly dissimilar paralogs, may indicate that the residue is important to maintain the three-dimensional fold involved in ligand or substrate binding, or protein-protein interactions.
Homology modeling assumes that the structure of a target protein can be solved based only on its primary amino acid sequence and its structural and evolutionary relatedness to a protein of known structure. Understanding the evolutionary relationships between template and target proteins, and the factors that drove their structural conservation, is extremely useful in homology modeling. Structure is usually more conserved than amino acid sequence, which is more conserved than nucleic acid sequence. One theory suggests that protein folds have evolved the robust ability to retain structure and function in spite of mutations (Taverna and Goldstein, 2002). Fragile folds that collapse in response to a few mutations might be selected against in favor of a robust protein fold which can evolve and adapt.
Proteins with 30% sequence identity, or 30 amino acids the same out of 100, will have similar folds (Sander and Schneider, 1991). Two sequences with greater than 25% sequence ID are considered highly related structures with true evolutionary homology, while those with less than 25% share some structural similarity and arguable homol-ogy (Sander and Schneider, 1991). Doolittle (1986) described this zone as the twilight zone, or a range of sequence identities that may be indicative of either divergent or convergent evolution. A sequence and structural analysis of proteins in the PDB found that structurally similar proteins could share sequence ID as little as 7-8%. Random amino acid sequences share about 4% sequence identity, and therefore the percentage of anchor residues, or those strictly required for structural relatedness, is actually only 3-4% (Rost, 1997).
A striking example of this statistic comes from the crystal structures of E. coli ribose- and lysine-binding proteins, which share the same fold despite little sequence identity (Kang et al., 1992). Surprisingly, the majority of related homologous structures in the PDB share less than 45% sequence ID (Rost, 1997). Amino acid mutations have been speculated to occur in intrinsically disordered regions, or loops that have little tendency for secondary structure, and have therefore evolved to allow the retention of structure and function. This theory was proven wrong by simulations which showed that, on the contrary, secondary structural elements can be maintained despite mutation accumulations, and in fact mutations in IDPs were much more likely to introduce secondary structure where previously there was none (Schaefer et al., 2010).
Inside of secondary structural elements, genetic drift appears to accumulate mutations in solvent-exposed regions with little functional value. A survey of the mutation rate for all amino acid types found that planar hydrophobic residues are the most conserved, followed by aliphatic residues. Charged residues were the least conserved residue type (Bowie et al., 1990). Some proteins may fold by a mechanism called hydropho-bic collapse, where hydrophobic residues nucleate the folding of a protein after or during translation by associating with each other and shielding themselves from water, and thereby shifting charged residues towards the outside (Nolting et al., 1995; Eaton et al., 1996). This process may explain why hydrophobic residues are well conserved.
Solvent-exposed residues are likely less well conserved unless they contribute to functional sites such as interaction interfaces. Sequence and structural conservation at protein-protein interaction interfaces is high. Histone proteins show greater than 98% sequence ID between humans and plants. Histones make ordered contacts with other histones and DNA itself, and thus there is high selection pressure on solvent-exposed residues. Lac repressor has two areas on its solvent-exposed surface that participate in interactions with the lac operator and inducer. These areas are conserved among members of the lac family, with little conservation elsewhere (Kisters-Woike et al., 2000). Conserved patches of solvent-exposed residues can indicate protein interaction domains, and this fact has been exploited by a program called consurf, which can be used to predict interaction interfaces based on a carefully constructed phylogenetic tree, homology models, and a multiple sequence alignment. Interaction domains must evolve reciprocal surfaces in order to continue interacting (Landau et al., 2005). Selection pressure increases as the number of binding partners utilizing the same domain increases more than one (Goh et al., 2000; Kisters Woike 2000).
Analysis of Protein-Ligand Interactions
Small-molecule ligands usually bind in pockets (Kuntz et al., 1982; Lewis, 1991). Ligand functional surfaces are often complementary to their binding space in terms of electrostatics and geometric shape (Altschul et al., 1997). These surfaces are frequently rough in order to fit a large amount of surface area and potential hydrophobic contacts into a defined amount of space (Pettit and Bowie, 1999). Algorithms have been developed to find concave surfaces as potential ligand-binding pockets (Kuntz et al., 1982; Peters et al., 1996). Given the genetic diversity of insects, comparative homology modeling, or comparing the same protein from many different organisms, is a great tool to find ligand-binding pockets.
Cytochrome C: A Case Study
Taxonomists routinely use the protein cytochrome C for DNA bar-coding and species identification because its amino acid sequence tends to be highly conserved among related species, with little variation between members of the same species (Hebert et al., 2003). Why is cytochrome c so conserved? The answer may partially lie in its size, the requirement for a heme-binding pocket, and its role as an interacting partner of proteins involved in both electron transport and apoptosis. As an electron transport protein, it binds a heme group, which can be oxidized or reduced to facilitate electron movement. Despite high sequence conservation, chimpanzee mitochondrial cytochrome oxidase systems suffer a 20% reduction in respiration capacity when introduced into human cell lines (Bar-rientos et al., 1998). This suggests that the evolution of reciprocal protein interaction interfaces between nuclear and mitochondrial proteins is required. The large number of interacting partners may place conservative selection pressure on these solvent-exposed residues. In the cyto-chrome c core, 22 of 103 amino acids are implicated in direct heme binding and/or required for the shape and hydrophobicity of the heme pocket and the overall fold. These 22 residues are highly conserved. Two more residues are solvent-exposed charged residues that may participate in partner binding and orientation (Takano and Dickerson, 1981).
Selecting a Template Structure
One easy method for template selection is performing a PSI-BLAST search against the RCSB Protein Data Bank from the NCBI blast homepage. Position Specific Iterative Blast uses a position-specific score matrix derived from the query for sequence comparison against the database of interest. PSI-BLAST can pick up weaker evolutionary relationships, and can give equal weight to the different domains of a protein instead of reporting the stronger more numerous relationships for one domain. PSI-BLAST works by first performing a regular protein blast, and then creating a multiple sequence alignment on the blast data, which are then used to create the position specific score matrix (Altschul et al., 1997; Schaffer et al., 1999). Another convenient feature of PSI-BLAST searches from NCBI is the option to view conserved domains using the conserved domain detection algorithm (CDD). CDD employs Reverse Position Specific Iterated Blast, or Reverse PSI-BLAST or RPS-BLAST (Marchler-Bauer et al., 2002). The two algorithms differ in the derivation of the position-specific score matrix from the database in RPS-BLAST and not from the query in PSI-BLAST (Schaffer, 1999). In the case of large multi-domain proteins it may not be necessary or even possible to model a whole protein due to little sequence conservation in intradomain regions. Some domains are known to fold and function independently of each other, and therefore it may not be necessary to model an entire protein.
Target-Template Sequence Alignment
Correct template-target sequence alignment is a critical factor in model quality. With greater than ~ 50% sequence ID, almost any algorithm will produce a suitable alignment (Rost, 1997) and thereby improve model accuracy. Alignment gaps are detrimental to the modeling process, and placing them in divergent or loop regions can improve model quality. The salign command in Modeller makes use of these two features, as well as placing gaps in solvent-exposed residues (Marti-Renom et al., 2000; Sali, 1995).
Modeling Suite Choice
When choosing a homology modeling software suite, the user should consider the suite’s accuracy and ease of use, and the algorithm employed. Target-template pairs with greater than 40% sequence identity produce similar structures regardless of the prediction server used. Modeling suites allow users more precise control over the modeling process, but often require knowledge of scripting languages. Users without sophisticated computer knowledge may want to choose packages with in-depth documentation and user support communities.
As sequence identity approaches the "twilight zone," modeling suite accuracy becomes more important. Servers such as I-Tasser (http://zhanglab.ccmb.med.umich.edu/I-TASSER/ [Zhang, 2008]) and Robetta (http://robetta.bakerlab.org [Kim et al., 2004]), and the Modeller suite (http://www.salilab.org/modeller/ [Sali, 1995]), use an approach to backbone generation that places restraints on values of the model structure. Backbone bond length, and PHI PSI and OMEGA angles, are constricted so that they can fall within a range of values derived from the template structure and a database of sequence structure relationships, also called a probability function. Modeller uses conjugate gradient optimization, beginning with local restraints and extending to global restraints, to optimize Newtonian force. Information on commonly used modeling programs is included in Table 5.
Critical Assessment of Protein Structure
Critical Assessment of Protein Structure (CASP) (http:// predictioncenter.org/casp8/groups_analysis.cgi) ranks the performance of prediction algorithms for completely automated servers. Some structural biologists choose to submit their experimentally determined structures for assessment in the contest prior to publication. Contestants are given the amino acid sequence of the target protein, and structure predictions are then made by either a human or server. The resulting structure files are compared to the previously determined structure by a number of algorithms, such as Dali (Holm and Rosenstrom, 2010) and Mammoth (Ortiz et al., 2002; Lupyan et al., 2005), which attempt to align the alpha carbon backbone or side chains and then determine the root mean square deviation (RMSD), or a derivative of RMSD, using the three difference dimensional coordinates of each structure file. Comparing two structure files can be somewhat subjective, and thus a number of alignment algorithms are employed. Alignment algorithms, and the databases of protein families that are often created with them, are useful for comparing models against other members to observe evolutionary traits. Protein family structures are also used in the beginning steps of modeling. After finding a suitable template, this structure can be compared to other members of the family.
Table 5 Commonly Used Modeling Programs
|
|
|
Server/user configured |
|
|
|
|
Name |
Website URL |
Citation |
Notes |
CASP ranking |
|
|
Modeller |
http://www.salilab.org/modeller/ |
Sali, 1995 |
User configured Python scripts |
User control is very high; great documentation and user-supported community |
N/A |
|
Itasser (Zhang Server) |
http://zhanglab.ccmb. med.umich.edu/I-TASSER/ |
Zhang, 2008 |
Automated server |
Threading approach allows structure predictions when template alignments are weak or non-existent |
Highest server ranking in CASP 8; 5th overall |
|
Robetta (BAKER-ROBETTA) |
http://robetta.bakerlab.org/ |
Kim et al., 2004 |
Automated server |
Comparative and de novo modeling |
2nd highest server rank in CASP 8; 22nd overall |
|
PDFAMS |
http://pd-fams.com/ |
Terashi et al., 2007 |
User configured scripts, some automation available |
Powerful; some software may need to be purchased |
4th overall CASP 8 |
|
Swiss Model |
http://swissmodel.expasy. org/workspace/ |
Bordoli et al., 2009; Kiefer et al., 2009 |
Automated server |
Accessed via a user-friendly Web Workspace or Deepview (Swiss-PDB-Viewer), a program available in the Microsoft Windows OS |
N/A |
Structural Determination
X-ray crystallography and nuclear magnetic resonance (NMR) imaging are the two primary methods of structure determination. X-ray crystallography can be used on much larger proteins with much better resolution. Some proteins cannot be expressed in sufficient levels and purified to a level amenable to either crystallography or NMR imaging. Crystallography has the drawbacks that some proteins will not crystallize, and in some cases the structure may actually be modified, or stuck in a single conformation that is not necessarily indicative of the dynamic conformational shifts the protein undergoes.
NMR, on the other hand, can be used to capture many types of motion. Backbone amide shifts have been used to determine ligand binding. Deuterium exchange experiments can reveal the change in solvent accessibility of particular functional groups. Proteins are grown in media containing hydrogen, and NMR recordings are performed in a solution of deuterium-labeled H2O. Hydrogen-deuterium exchange events can then be monitored. Another advantage of NMR is that structures are not modified by the crystallization process, and are viewed in a more natural aqueous environment. However, not all proteins are easily soluble in solution.
Metabolomics
Metabolomics involves the high-throughput characterization of all small-molecule metabolites and the products of biochemical pathways. The responses of biological systems to genetic or environmental changes are often reflected in their metabolic profiles. There are three major categories in metabolomics. The first is targeted metabolomics, which documents changes in metabolites in response to environmental conditions the insects encounter. The second, metabolic profiling, qualitatively and quantitatively evaluates metabolic collections. The third, metabolic profiling, collects and analyzes data from crude extracts to classify them based on all metabolites rather than separating them into individual metabolites. Gas chromatography and LC-MS are used for the identification and quantitation of metabolites. Nuclear magnetic resonance methods are employed for de novo identification of unknown metabolites. In insects, metabolomics could help in classification, studies on toxicology of insecticides, and safety testing of insecticides, and to monitor effects of genetic and environmental conditions on insect physiological processes.
Systems Biology
As stated earlier, systems biology takes a holistic view of a system or process by attempting to integrate all the data generated by various independent pathways technologies, and analyzing them together to formulate a hypothesis or model. Researchers working on insects have just begun to apply the systems biology approach to achieve an integrated view on the functioning of insect physiological systems. One such example is the recent study on D. melanogasterphagasome. Upon encountering microbes or other antigens, phagocytes internalize these particles into phagosomes to initiate destruction of these immune agents. Stuart and colleagues applied the systems biology approach to address the complex dynamic interactions between proteins present in the phagosomes and their involvement in particle engulfment (Stuart et al., 2007). This analysis identified 617 proteins associated with D. melanogaster phagosomes. The 617 phagosome proteins were used to prepare a detailed protein-protein interaction network, and 214 of the 617 phagosome proteins were mapped to a protein-protein interaction network. RNA interference was then employed to determine the contribution of each protein in microbe internalization. RNA interference studies identified gene coding for proteins that are known to function in phagocytosis. In addition, these studies also identified novel regulators of phagocytosis. These pioneering systems biology studies have provided new insights into functional organization of phagosomes. Such holistic approaches applied to various physiological systems in insects may lead to better understanding of the functioning of these systems.
Conclusions and Future Prospects
The rapid development of next generation sequencing (NGS) technologies during the past four years, following the domination of the automated Sanger sequencing method for almost two decades, could revolutionize the way of thinking about scientific approaches in insect research. The impact of the introduction of NGS technologies into the market is similar to the early days of PCR, with imagination being the only limiting factor for their use. It will be possible to sequence genomes of insects at $1000/genome in the not too distant future. The availability of genome sequences of almost every insect species of interest will help with research in every field of entomology. Advances in omics fields, as well as both forward and reverse genetics and RNA interference approaches, will also help in advances in research on insects. In the near future, molecular phylogenetics studies will use whole-genome sequences for insect taxonomy. Neurobiologists and physiologists will use systems biology approaches to understand the complexity of neuronal signaling and other physiological processes.