1. X chromosome inactivation
1.1. The mammalian sex chromosomes
Dioecy, the presence of two distinct sexes, is common amongst animals. In mammals, sex determination is controlled by distinct sex chromosomes (the X and Y), which are believed to have originated from an autosomal chromosome pair by the acquisition of a major sex-determining locus and subsequent divergence (Ayling and Griffin, 2002). Suppression of recombination near the sex-determining locus would be evolutionarily favored; however, the absence of exchange between the homologs would also lead to the degeneration of the sex-determining Y over evolutionary time (Ohno, 1967). The human X and Y maintain identity in only a small region of homology at the tips of both the long and short arms, referred to as the “pseudoautosomal” region (PAR), which undergoes recombination during male meiosis. The male-specific region of the Y also retains 20 genes or gene families derived from X chromosomal sequences in addition to testis-specific multicopy sequences, many of which are arrayed in palindromic segmental duplications (Skaletsky et al., 2003). The presence of such repeats has been proposed as a mechanism allowing gene conversion to protect haploid Y genes against their inevitable decay due to acquisition of mutations (Rozen et al., 2003). The X retains much of its old autosomal information, including numerous important housekeeping genes, despite being a relatively gene-poor chromosome containing approximately 5% of the haploid genome but only about 3% of the genes (Graves et al., 2002).
1.2. Dosage compensation in mammals and other species
In response to the degeneration of the Y, there would have been adaptive pressure to increase expression from the single expressed X-linked copy of genes. Indeed, dosage of X-linked genes, despite their monoallelic expression, seems comparable to autosomal genes, as determined by comparison of expression levels of a variably X-linked gene in different species (Adler et al., 1997), microarray analysis (Disteche et al., 2001), or impact of aneuploidy (Mizuno et al., 2002). In addition, excessive X-linked gene expression in females relative to males established the need for dosage compensation, which is achieved by X chromosome inactivation, whereby a single X is inactivated in female cells early in mammalian development. In individuals with an abnormal number of X’s (e.g., 45,X; 47,XXX; 47,XXY, etc.), a single X remains active no matter how many are present (the “n-1″ rule). In triploids, two X’s can be active, implicating autosomal copy number in determining the number of X’s that remain active (Migeon, 2002).
Dosage compensation also occurs in other dioecous organisms, utilizing a variety of mechanisms. In the fruit fly, Drosophila melanogaster (males XY, females XX), hypertranscription of most genes along the single male X matches the output of the female’s two X’s, and in the nematode, Caenorhabditis elegans (males XO, hermaphrodites XX), dosage compensation is accomplished in hermaphrodites by downregulation of X-linked gene transcription from both X’s (reviewed in Marin et al., 2000). In birds (males ZZ, females ZW), there is some evidence for dosage compensation; however, it appears that this is not accomplished by the silencing of one Z chromosome as biallelic expression in the male is observed (Kuroiwa et al., 2002). Despite their substantial differences, dosage compensation in mammals, flies, and worms is thought to involve the control of chromatin structure, and in flies and mammals, it apparently involves functional RNAs (XIST in mammals, see below; and roX in flies (Meller et al., 1997; see also Article 27, Noncoding RNAs in mammals, Volume 3)). Mammalian X chromosome inactivation is unique amongst these mechanisms in that it requires differentiation between an active and inactive X within the same nuclear environment.
2. Features of the inactive X
The inactive X (Xi), as befits facultative heterochromatin, shares general features with constitutive heterochromatin. These include suppression of gene expression, condensation of chromatin (forming the Barr body), late replication, nuclease insensitivity, hypoacetylation of histones, and hypermethylation of DNA. However, several unique features characterize heterochromatin of the Xi, including the association of a large untranslated RNA, the X-inactive specific transcript (XIST in human or Xist in mice), and a nonrandom distribution of variants of the core histone H2A. Once the inactive state is established, silencing is faithfully maintained throughout subsequent mitotic cell divisions, resulting in clonal populations, that, in the case of some tumors, could include 109 cells (Linder and Gartler, 1965). Thus, X inactivation is a paradigm for both the establishment of a differentiated chromatin configuration and cellular memory of that differentiated state. Figure 1 summarizes some of these features of the Xi, which are discussed below.
2.1. Gene silencing
Genes that continue to be shared between the X and Y, notably those in the PARs, would have no need for dosage compensation, as they would continue to be present in two copies in both males and females, and thus it was anticipated that some genes would “escape” inactivation and continue to be expressed from the Xi (Lyon, 1962). What is surprising, however, is the number of genes escaping inactivation in humans. Over 15% of human genes continue to be expressed from the inactive X chromosome (Carrel et al., 1999), and clearly, not all of these have Y homologs. Thus, a dosage imbalance may continue to exist, whereby human females express more X-linked gene product than males. Interestingly, in mice, fewer genes have been identified to escape inactivation (reviewed in Disteche et al., 2002). Genes that escape inactivation are not all “fully” expressed from the Xi, and several human genes have been shown to have variable inactivation, escaping inactivation in some females but not in others (e.g., TIMP1 and CHM1 (Anderson and Brown, 1999; Carrel and Willard, 1999)). The question of how some genes on the chromosome continue to be expressed while others are being silenced by a cis-acting mechanism is a fascinating biological question, and may also have clinical implications for disease, as discussed below. For many of the genes that have been examined, the general features of silent chromatin (e.g., replication timing, DNA methylation, and histone modifications) are not associated with an active gene on the Xi (see Disteche et al., 2002 for a review). A clue to the mechanism by which genes escape silencing may lie in the fact that many, although not all, of the genes expressed from the Xi are located in clusters, implicating a regional mechanism in their failure to be silenced (e.g. Tsuchiya and Willard, 2000). It is not clear whether expression from the Xi results from an initial inability to undergo inactivation or instead reflects an inability to maintain the silent state. The mouse Smcx gene is subject to inactivation in a proportion of embryonic cells; however, reactivation of the gene occurs very early in development (Lingenfelter et al., 1998). Such an early reactivation could be difficult to distinguish from the failure of inactivation spreading. Given the large number of genes escaping inactivation in humans, it is likely that there is more than a single mechanism by which expression from the Xi is acquired.
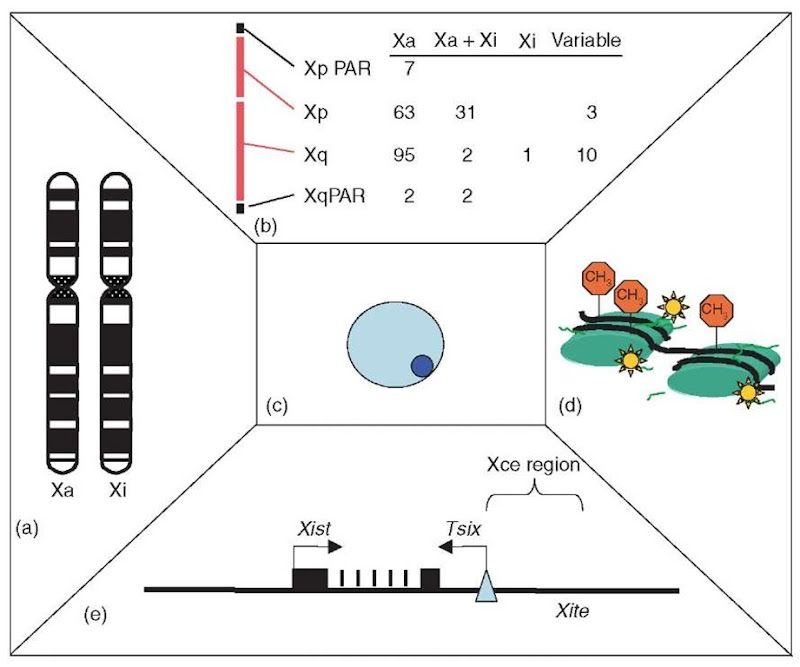
Figure 1 Facets of X inactivation. (a) In mammals, dosage compensation must differentiate between the active and inactive X chromosome in the same nuclear environment. Shown here are ideograms of the human X chromosomes. (b) Furthermore, in humans, approximately 15% of genes escape inactivation, being expressed from the Xa and Xi. As tabulated in this drawing, the majority of genes escaping inactivation are located on the short arm of the X (data from Carrel L, Cottle AA, Goglin KC and Willard HF (1999) Proceedings of the National Academy of Sciences of the United States of America, 96, 14440-14444). (c) In interphase nuclei, the Xi can manifest as the Barr body, which stains darkly with DAPI, as diagrammed in this cartoon. (d) Modifications of the chromatin on the Xi include methylation of the DNA at the promoters of X-linked genes (shown as red stop signs) and hypoacetylation of histones as well as hypermethylation at lysine 9 and 27 of histone H3 (represented as stars). (e) The Xist gene is necessary for the establishment of inactivation, and in mice, Xist expression is regulated by the antisense Tsix transcript that initiates transcription at a CpG island (blue triangle) shown to bind CTCF. The choice of X to inactivate is influenced by the Xce locus and the recently defined Xite region.
2.2. Barr body formation
During interphase, the human Xi forms a darkly staining body, the Barr body, which is seen associated with the nucleolus (Barr and Bertram, 1949) or at the nuclear periphery (Dyer et al., 1989), the usual sites for heterochromatin. Three-dimensional reconstructions of the Xi territory suggest that rather than being substantially more condensed than the active X (Xa), the principal difference between the two chromosomes is in shape (Eils et al., 1996). Thus, the intense staining of the Barr body with DNA dyes likely reflects the distinctive packaging of DNA on the Xi; however, to date, only a limited number of proteins have been shown to associate specifically with the Xi.
The Xi is characterized by the accumulation of variant histone H2A isoforms termed macrohistone H2A1 and 2 (Costanzi and Pehrson, 1998; Chadwick and Willard, 2001). These proteins are very similar to H2A, but include a large carboxy-terminal domain, and their enrichment at the Xi forms a characteristic structure in the female nucleus, referred to as a macrochromatin body (MCB). MacroH2A has been shown to have a transcriptional repressive effect (Perche et al., 2000), potentially caused by both interfering with transcription factor binding and chromatin remodeling (Angelov et al., 2003). Deletion of Xist after inactivation results in loss of macroH2A association with the Xi (Csankovszki et al., 1999), while induction of Xist is sufficient to induce MCB formation (Rasmussen et al., 2001), suggesting a direct or indirect association between the Xist RNA and macroH2A. Such an association is further supported by coimmunoprecipitation of Xist by macroH2A antisera (Gilbert et al., 2000).
Chadwick and Willard (2003b) recently examined a variety of chromatin proteins for enrichment on the Xi and demonstrated elevated levels of heterochromatin protein-1 (HP1), histone H1, and the high mobility group protein HMG-I/Y at the territory of the Xi in interphase human cells. The association, or at least the visualization of the association, of these various proteins with the Xi seems to be cell-cycle and perhaps cell-type specific, and often correlates with visualization of the Barr body (Chadwick and Willard, 2003a). Surprisingly, another protein shown to be associated with the inactive X and required for XIST localization is the BRCA1 protein (Ganesan et al., 2002), which has multiple cellular functions, including chromatin remodeling (Bochar et al., 2000). Furthermore, transient association of the polycomb group complex proteins Eed and Enx1 is seen in the earliest stages of X inactivation (Wang et al., 2001; Mak et al., 2002; Silva et al., 2003). Interestingly, these proteins are also required early in development for the monoallelic expression of some imprinted loci (Mager et al., 2003). Monoallelically expressed genes (both imprinted and X-linked) seem to have dimethylation of histone H3 at lysine (K) 4 restricted to the promoter and not the body of the gene, a mark that is already present in totipotent embryonic stem (ES) cells (Rougeulle et al., 2003). Intriguingly, this mark is not seen for the Smcx gene that shows rapid reactivation from the Xi.
2.3. Replication timing
Synchrony of replication origin firing in mammalian cells results in megabase-sized domains that replicate at a similar time during S phase (see Goren and Cedar, 2003 for a review). In general, expressed genes tend to replicate earlier in S phase than silenced genes (Woodfine et al., 2004). Late replication of the Xi is observed after pulse labeling of cells with bromodeoxy-uridine, and while there is some cell and tissue specificity to the replication patterns (e.g., Willard, 1977), there are regions of the inactive X that replicate earlier than the rest of the chromosome (e.g., XpPAR), and thus have been hypothesized to contain genes that do not undergo inactivation (Schempp and Meer, 1983). Assessment of replication timing for individual genes supports this suggestion as genes escaping inactivation show synchronous DNA replication between the Xa and Xi (e.g., Boggs and Chinault, 1994; Hansen et al., 1996). The origins of replication for the HPRT and G6PD loci are the same on the Xa and Xi (Cohen et al., 2003), implicating the yet-to-be-elucidated epigenetic determinants that delay replication firing on the Xi. A long-standing question is whether replication timing dictates the structure of chromatin or vice versa, and recent DNA injection experiments suggest that timing of replication is sufficient to alter transcriptional activity (Zhang et al., 2002). However, the transition from synchronous to asynchronous replication of the two X’s in early embryos appears to follow alterations in histone modifications, suggesting that chromatin changes may establish the late replication of the Xi (Chaumeil et al., 2002).
2.4. Histone modifications
In addition to variants of the core histones, chromatin can vary in the modifications of histones (see Article 27, The histone code and epigenetic inheritance, Volume 1). Many amino acid residues of core histones, located predominantly within the tail regions, have the potential to acquire a range of covalent modifications, including acetylation, methylation, ubiquitinylation, and phosphorylation. It has been hypothesized that a specific combination of modifications confers a particular property to the local chromatin, referred to as the “histone code” (Jenuwein and Allis, 2001), and the Xi shows modifications common to other regions of heterochromatin. Specifically, the Xi shows hypomethylation at K4 and 36, and arginine (R) 17 of histone H3; hypermethylation at K9 and K27 of histone H3; hypoacetylation at R2, R12, and R26 of histone H3 and K5, K8, K12, and K16 of histone H4 (see Chaumeil et al., 2002; Chadwick and Willard, 2003a). How the histone modifications associated with the Xi determine or influence interactions with other chromatin proteins that are involved in X inactivation is largely unknown, however, acquisition of H3 modifications is one of the first detectable events after Xist RNA coating of the presumptive Xi (Chaumeil et al., 2002). The Suv39h histone methyltransferase responsible for constitutive heterochromatin histone K9 methylation does not seem to be necessary for Xi histone methyla-tion (Peters et al., 2002), while the Eed-Enx1 histone methyltransferase complex shows only a transient association with the Xi (Silva et al., 2003), implicating the involvement of additional chromatin-modifying enzymes in X inactivation.
2.5. DNA methylation
In mammals, symmetrical DNA methylation at CpG dinucleotides is a stable and clonally heritable epigenetic modification that is maintained by the action of a maintenance methyltransferase acting preferentially on hemimethylated substrates that result from DNA replication (Riggs, 1975). DNMT1 is generally ascribed this maintenance methylase activity, while DNMT3A and B are considered de novo methylases; however, understanding of how methylation patterns are established is limited (see Meehan, 2003 for a review). DNA methylation can regulate gene expression either by directly blocking transcription regulatory factors from binding to their target sequences or through several methyl-CpG-binding proteins that “read” DNA methylation patterns and form complexes with transcriptional repressor proteins including histone-modifying enzymes (Jones et al., 1998; Fuks et al., 2003). The CpG islands in the promoter regions of X-linked genes are heavily methylated on the Xi, in contrast to their unmethylated state on the Xa and the general state for autosomal genes. This epigenetic modification is one of the latest to be acquired during the process of X inactivation, following inactivation by several days and implicating methylation in maintenance rather than initiation of X-linked gene silencing (Keohane et al., 1996). X inactivation in marsupials and eutherian extraembryonic tissues, where there is reduced DNA methylation, is less stable (e.g., Kaslow and Migeon, 1987; Migeon et al., 1985). Similarly, patients with immunodeficiency-centromeric instability-facial anomalies syndrome (ICF, MIM 242860), which results from mutations in DNMT3b, have some hypomethylated CpG islands, and show reactivation of some X-linked loci in some cells (Hansen et al., 2000). Interestingly, they also show hypomethylation of LINE elements on only the Xi, suggesting that another methyltransferase is involved in LINE element methylation on the Xa and autosomes (Hansen, 2003). Furthermore, inhibition or loss of DNA methyltransferase leads to derepression of X-linked genes, providing experimental evidence for the importance of DNA methylation in maintenance of X inactivation (Venolia et al., 1982; Sado et al., 2000). Synergism between XIST localization, DNA methylation, and histone modifications results in extremely stable silencing of the Xi (Csankovszki et al., 2001).
3. Process of inactivation
X inactivation can be envisioned as a cycle of events. Reactivation of the entire Xi occurs normally during oogenesis, while the X chromosome is transcriptionally inactivated together with the Y chromosome in spermatogenic cells shortly before or during early meiotic prophase. Such male meiotic sex chromosome inactivation (MSCI) has been suggested to mask the nonsynapsed regions of the sex chromosomes, possibly in order to avoid the checkpoint that causes germ cell death in response to defective chromosome synapsis; prevent the initiation of recombination events in the nonsynapsed regions of the sex chromosomes; or ensure efficient sex chromosome synapsis (reviewed in Turner et al., 2002). Inactivation occurs in female somatic cells at approximately the time of implantation during the late blastula stage, and is observed first in the extraembryonic tissues, where it is imprinted in mice (see below, and Article 41, Initiation of X-chromosome inactivation, Volume 1). Recent results suggest that the mouse X may be partially inactivated at the 2-cell stage (Huynh and Lee, 2003), and that early imprinted silencing is reversed in cells of the inner cell mass prior to random inactivation (Okamoto et al., 2003). As human X inactivation is not imprinted, it is not known to what extent events in very early development will be similar between humans and mice.
Much of our current understanding about the molecular mechanisms involved in the regulation of X inactivation is derived from the studies of mice, particularly ES cells that can undergo inactivation upon differentiation. A time line of inactivation suggests that first XIST expression is stabilized, coating the future Xi, and then histone modifications, gene silencing, asynchronous replication, and macroH2A accumulations occur (Chaumeil et al., 2002). The mechanism by which inactivation spreads in cis along the 160-megabase chromosome remains unsolved (see Article 40, Spreading of X-chromosome inactivation, Volume 1); however, it has been hypothesized that there are “way-stations” or booster elements distributed along the chromosome that serve to assist in the propagation of inactivation (Riggs et al., 1985). In X/autosome translocations, silencing can spread into the autosomal region; however, spread within the autosome does not seem to be as effective as on the X (White et al., 1998). Thus, way-stations are likely concentrated on the X, and Lyon has hypothesized that LI elements, which are enriched on the X, could be the way-stations (Lyon, 1998). Several bioinformatic studies have examined the characteristics of regions that escape inactivation (Bailey et al., 2000; Friel et al., 2002). However, as mentioned above, such regions may escape inactivation either because of failure to respond to the initial signal or inability to maintain silencing; and thus, the enrichment of particular elements in these regions may reflect a role in either the spread or the maintenance of silencing.
3.1. The X inactivation center
A critical region necessary for inactivation of the X was defined by the analysis of the ability of rearranged X’s to undergo inactivation. This region, the X inactivation center (XIC), was refined to 1Mb of human Xq13 (Leppig et al., 1993), and more recent mouse transgenic studies suggest that the Xic is no more than 450-kb long (Lee et al., 1996). This region includes the XIST/Xist locus, as well as several cis-acting loci that influence the random choice of X to inactivate (reviewed in Rougeulle and Avner, 2003; and discussed below).
3.1.1. XIST
XIST/Xist, a large (>15kb) alternatively processed, noncoding RNA, is the only gene known to be transcribed from the Xi but not from the Xa in somatic cells, and the RNA associates in cis with the Xi as part of the Barr body (Brown et al., 1991; Brockdorff et al., 1991; Clemson et al., 1996). The Xist gene is required for inactivation, and transgenes of Xist are able to induce inactivation of autosomes, identifying Xist as the principal component of the Xic (see Rougeulle and Avner, 2003). Prior to the onset of inactivation, Xist is unstable and expressed at a low level from all X chromosomes. Upon differentiation, Xist RNA is stabilized and coats the presumptive Xi, while the low-level pinpoint of Xist RNA on the Xa is subsequently extinguished (Panning et al., 1997; Sheardown et al., 1997).
Xist is known to be essential for the initiation of X inactivation based on targeted deletions in the mouse. Specifically, an Xist deficiency in female ES cells abolishes X inactivation potential in cis, enabling inactivation only of the chromosome carrying the wild-type Xist allele. Subsequent experiments have suggested that the Xist RNA levels determine which X undergoes inactivation (Nesterova et al., 2003). The continued expression and association of the transcripts with the Xi in somatic cells also implicated the RNA in the maintenance of the Xi; however, deletion of XIST/Xist after inactivation does not eliminate gene silencing (Brown and Willard, 1994; Rack et al., 1994; Csankovszki et al., 1999), although it does eliminate macroH2A association and increases the rate of reactivation of individual genes (Csankovszki et al., 2001). Reactivation in the absence of Xist is synergistically enhanced by treatment with demethylating or deacetylase-blocking agents, emphasizing the highly redundant nature of the maintenance of inactivation (Csankovszki et al., 2001).
Xist by itself, however, is not sufficient to induce chromosome-wide gene silencing in rodent somatic cells where reactivation of the endogenous Xist or induction of Xist-encoding transgenes produce localized Xist that does not inactivate the X (Clemson et al., 1998; Hansen et al., 1998; Rasmussen et al., 2001). Nevertheless, in human-transformed cells, it has been demonstrated that XIST expression can induce hallmarks of silencing, suggesting that the ability of XIST/Xist to induce inactivation may be species or cell-type specific (Hall et al., 2002). Unexpectedly, in human/mouse hybrid cells, XIST is unable to localize to the human X, implicating species-specific factors in the chromatin recognition of XIST (Clemson et al., 1998; Hansen et al., 1998).
Xist RNA consists of separable domains for silencing and coating the chromosome in cis (Wutz et al., 2002). The transcriptional silencing activity of the transcript can be attributed to a domain at the 5′ end of the RNA that shows sequence conservation in all species in which Xist has been analyzed. An Xist RNA deleted for this 5′ domain coats, but does not silence, the X. Furthermore, only RNAs that were capable of coating the chromosome were functional in silencing. The separable motifs within Xist RNA probably recruit protein complexes to mediate coating and/or silencing of the X.
3.1.2. Tsix
A gene overlapping and antisense to Xist, named Tsix, has been identified in mouse ES cells; it is expressed from both X’s before and during, but not after, differentiation (Lee et al., 1999; Debrand et al., 1999). Tsix initiates at a major transcription start site 13 kb downstream of the Xist 3′ end and includes four exons that undergo alternative splicing (Mise et al., 1999; Shibata and Lee, 2003). Like Xist, Tsix has no open reading frames, and its RNA is found in the nucleus. Tsix expression seems opposed to Xist expression, and truncation of the Tsix transcript led to complete nonrandom inactivation of the targeted X chromosome (Luikenhuis et al., 2001). Induction of Tsix transcription during ES cell differentiation, on the other hand, caused the targeted chromosome always to be chosen as the active chromosome (Stavropoulos et al., 2001). These results suggest that Tsix is a critical negative regulator of Xist. Tsix knockout mice show a dramatic parent-of-origin phenotype consistent with Tsix being a central player in imprinted inactivation (Lee, 2000). Although an antisense reminiscent of Tsix has been reported in humans, the roles of the human TSIX remain controversial (Migeon et al., 2001; Chow et al., 2003).
3.1.3. Xce and random versus imprinted X chromosome inactivation
In eutherian embryonic cells, the choice between inactivation of the maternally or paternally derived X is random. However, in marsupials, the paternal X is inactivated in all cells (Cooper et al., 1971), and a similar imprinted inactivation is found in the extraembryonic tissues of mice in which the paternal X is always inactive (Takagi and Sasaki, 1975). Tsix has been suggested to be the cis-acting imprinting factor that protects the maternal X from silencing, thus playing a critical role in imprinted X inactivation within developing mouse extraembryonic tissues (Lee, 2000; Sado et al., 2001). Preferential paternal inactivation in the extraembryonic tissues is also observed in cows (Xue et al., 2002). While there have been some conflicting results, it appears that despite some limited bias toward paternal inactivation in early trophoblast, inactivation in the extraembryonic tissues of humans is not imprinted (see review by Migeon, 2002).
In mice, the X controlling element (Xce), biases inactivation toward one X, with 20-30% distortions from random inactivation being observed in mice with different alleles of the Xce locus (Cattanach and Issacson, 1967). Xce maps 3′ to Xist (Simmler et al., 1993), and the Xite locus has been proposed as a candidate for Xce activity. Xite is a region of intergenic transcription beyond Tsix that is associated with DNaseI hypersensitivity (Ogawa and Lee, 2003). A human equivalent to Xce has not been identified, although a screen of 38 families identified several families with a statistically significant skewing of X inactivation (Naumova et al., 1998). Mutations in the XIST promoter have been associated with deviations in X inactivation, but this strong skewing of inactivation was not consistent throughout families (Plenge et al., 1997; Tomkins et al., 2002). A mutational screen for modifiers of the X inactivation ratio in mouse identified two autosomal dominant loci, which have yet to be characterized (Percec et al., 2002).
4. Clinical consequences and considerations
X inactivation results in mammalian females being mosaics, composed of cell lines in which either the maternally or the paternally inherited X is inactivated. This mosaicism is generally sufficient to protect females from X-linked disease, but females may show “patchy” expression if the gene product is cell restricted, or disease, depending upon the randomness of their X inactivation pattern. There are also X-linked dominant disorders, many of which are believed to be male lethal (e.g., Rett syndrome, MIM 312750 or incontinentia pigmenti, MIM 308300), although others seem to manifest predominantly in females, possibly due to metabolic interference (e.g., craniofrontonasal syndrome MIM 304110 (Wieland et al., 2002)).
The process of inactivation also minimizes the impact of X aneuploidy, as all but one X will be inactivated. However, the lack of an Xi in females results in Turner syndrome, demonstrating that the Xi provides critical gene products for normal development, which are presumably provided by the Y in males. Female mice lacking an Xi show less phenotypic consequence, likely reflecting the fact that fewer genes are expressed from the Xi in mice (Disteche et al., 2002). The X/Y homologous genes represent only a small proportion of the genes that are expressed from the Xi in humans (Carrel et al., 1999), and thus, there is the possibility for considerable differential expression between males and females. The role of differential X-linked gene expression is quite difficult to separate from the substantial hormone-induced differences between males and females; however, there is a developing literature supporting difference in sex-chromosome gene expression between the sexes, particularly in brain (e.g., Xu et al., 2002; Good et al., 2003).
Deviations from the normal random inactivation pattern can occur either owing to primary influences upon the choice of X to inactivate or more likely owing to postinactivation selection for or against a population of cells with one X active (Migeon, 1998). A large number of assays have been developed that combine an X-linked polymorphism with a distinguishable feature of the Xi or Xa (generally methylation or expression) to allow determination of the extent of skewing of inactivation. In addition to being used to determine skewing in females suspected of carrying X-linked defects, these assays are often used for the analysis of clonality in tumor tissue. If a tumor originates from a single cell, tumor tissues will have the same inactivation pattern in all cells. This nonrandom pattern of inactivation has been observed in a wide range of neoplastic tissue, demonstrating the monoclonal origin of the neoplasia (reviewed in Busque and Gilliland, 1998).
Although the analysis of inactivation patterns in females has been valuable in studies of carriers of X-linked disease and for determining the clonality of tumors, our limited understanding of the contribution of different causes of skewing confounds the utility of such studies. Skewed inactivation patterns have been reported in association with females with breast cancer, ovarian cancer, or females with recurrent spontaneous abortions, and these associations have yet to be satisfactorily explained (Buller et al., 1999; Kristiansen et al., 2002; Sangha et al., 1999, Lanasa et al., 1999, Uehara et al., 2001). One possibility is that X-linked mutations underlie such associations (e.g., Lanasa et al., 2001). Alternatively, skewing of inactivation may reflect a limitation in the number of viable cells that contribute to an embryo, as is the case for the skewed inactivation frequently observed in fetuses or newborns associated with confined placental mosaicism, in which a trisomic conceptus likely underwent a rescue event leading to a limited population of disomic cells (Lau et al., 1997). Thus, further studies are necessary for the determination of the cause or causes of these associations.