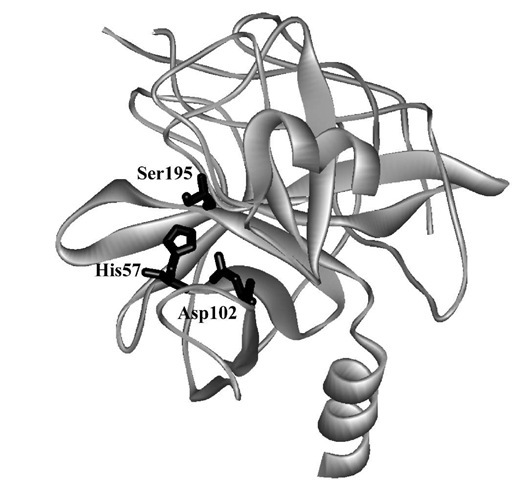
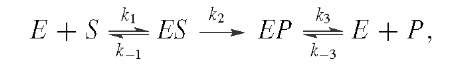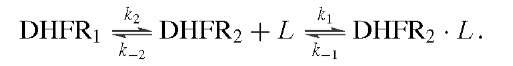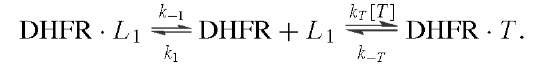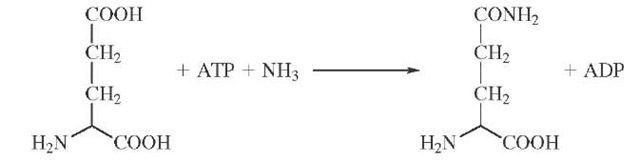Alpha-chymotrypsin (Fig. 2) catalyzes the facile hydrolysis of peptide bonds, in particular those adjacent to the carboxyl group of aromatic amino acids (tryptophan, ty-rosine, phenylalanine) as well as a variety of esters derived from similar N-acylated amino acids. The enzyme has been the subject of intensive mechanistic study, most of which occurred well before a crystal structure was available.
where the rate constants are in the order![]()
The amplitude of the burst can provide the concentration of active sites in an enzyme preparation. By varying the concentration of S, one can find values for![]() and k3. There are many variations on transient kinetics,as will be illustrated in our case studies of individual enzymes.
and k3. There are many variations on transient kinetics,as will be illustrated in our case studies of individual enzymes.
A key insight was provided by studying the enzyme-catalyzed hydrolysis of ^-nitrophenyl acetate. Transient kinetic studies revealed burst kinetics (Fig. 3) with an initial rapid liberation of ^-nitrophenolate followed by a slower steady-state rate. The biphasic time course is consistent with the existence of two intermediates (ES and acyl-E), with the second accumulating owing to its slower breakdown to product. The intermediate is a covalent enzyme species acylated at serine-195 (see Fig. 2), a fact initially revealed by chemically esterifying this enzyme residue specifically and irreversibly with diisopropylphos-phorofluoridate. No burst kinetics is seen with amide substrates because the acylation step limits turnover. The same intermediate, however, is formed as shown by partitioning experiments in which an exogenous nucleophile such as hydroxylamine is added to compete with water in the deacylation step. The result revealed equivalent levels of hydroxamate and acid products formed from either amide or ester substrates derived from a common amino acid, which implicated the presence of the intermediate in both enzyme-catalyzed processes.
FIGURE 2 The crystal structure of a-chymotrypsin showing the catalytic triad of amino acid side chains.
FIGURE 3 Plot of the burst in hydrolysis of p-nitrophenyl acetate. The concentration of product is observed as a function of time.
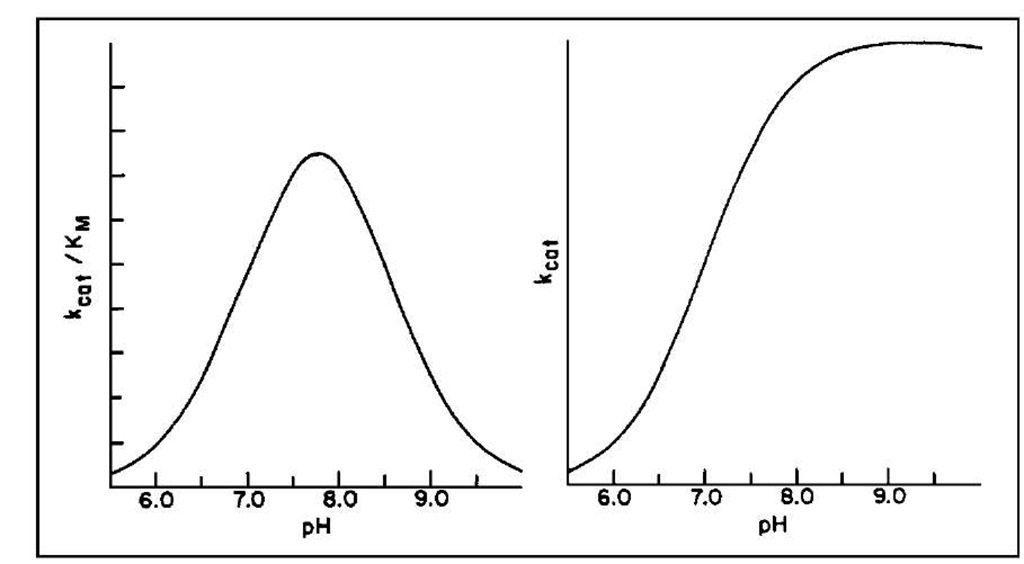
The pH dependence of the steady-state kinetic parameters is shown in Fig. 4 and implicates the ionization of two groups in the free enzyme and one in the ES complex. These data combined again with chemical modification studies (now superseded by site-specific mutagenesis) implicated histidine-57 (pKa ~ 7) and the N-terminal amino acid isoleucine (pKa ~ 8.5). The latter forms a salt bridge with aspartate-194 that helps maintain the active structure of the enzyme; the former is involved in general acid-base chemistry at the active site.
These data, along with further information derived from the reaction of specific substrates with the enzyme by using stopped-flow methods, led to the elucidation of a kinetic sequence that consistently implicated the acyla-tion and deacylation of Ser195 assisted by His57 and Asp102. The crystal structure of chymotrypsin (Fig. 2) reveals that these three residues form a catalytic triad, a feature repeated for many hydrolytic enzymes. This triad operates within a well-defined binding site that is lined with nonpolar amino acids capable of van der Waals interactions with polypeptide substrates containing aromatic side chains. A plausible mechanism is outlined in Fig. 5 in terms of the chemistry occurring during the individual kinetic steps.
The key features of this mechanism require the participation of the serine hydroxyl as a nucleophile whose attack on the carbonyl of the substrate is facilitated through proton abstraction by the imidazole nitrogen of His57 and its redonation to the amine-leaving group. Deacylation of the enzyme follows general base catalysis of water attack again by His57 and the return of the enzyme to its resting state. Catalysis of the chemical process through the participation of the side chains of an enzyme in proton, hydride, and electron transfer is a hallmark of enzyme catalysis and can occur efficiently in the confines of the active site owing to the optimal alignment and juxtapositioning of the substrate for chemical reaction.
B. Dihydrofolate Reductase
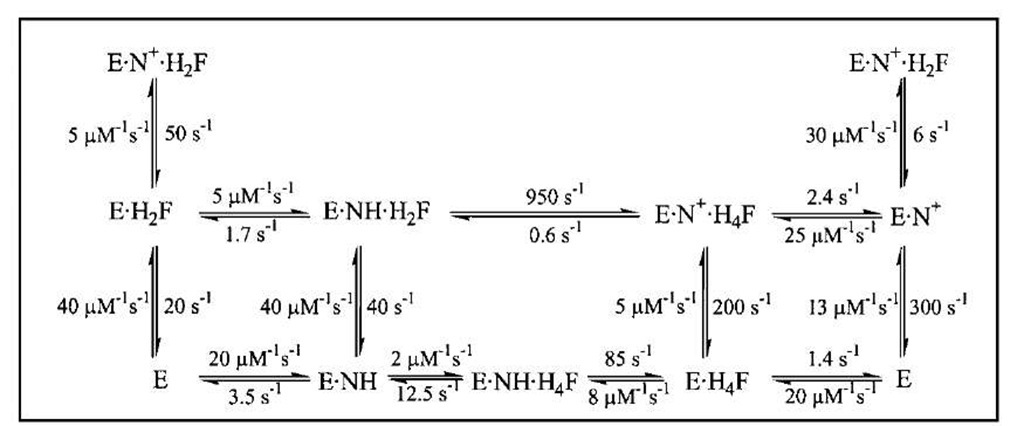
Dihydrofolate reductase (DHFR) catalyzes the reduction of 7,8-dihydrofolate (H2F) by nicotinamide adenine din-ucleotide phosphate (reduced form) (NADPH) to form 5,6,7,8-tetrahydrofolate (H4F), a key step in furnishing the parental cofactor needed for de novo pyrimidine and purine biosynthesis. The enzyme has been the target of antitumor and antimicrobial drugs. A complete kinetic scheme (Fig. 6) obtained primarily through transient kinetics has been described for the enzyme from Escherichia coli as well as other sources and provides a second case study as to how to define the catalytic process.
FIGURE 4 The pH dependence of Abat/Kvi and kcat for the a-chymotrypsin-catalyzed hydrolysis of esters and amides.
Measurement of the rates of binding and dissociation of substrate and cofactors provided valuable insights into the identity of rate-limiting kinetic steps in the scheme shown in Fig. 6. Two procedures were used. In the first, direct observation of changes in the intrinsic enzyme or NADPH fluorescence upon ligand binding showed that the addition of ligand was biphasic in accord with the existence of two conformers, of which only one bound the ligand:
The rate of the initial fast phase and its amplitude are associated with the binding of L to DHFR2 (ki, k_i) and the level of DHFR2; the rate of the second phase is the conversion of DHFR1 to DHFR2 (k2). The method was extended to the binding of a second ligand to binary DHFR2 L complexes and revealed that the binding of various ligands was near the diffusion-controlled limit.
In the second procedure a competitive trapping technique was employed in which the enzyme-ligand complex is mixed with an excess of a second ligand that competes for the binding site. With this method, k_1 is measured accurately when![]() This
This
procedure identified a preferred pathway for dissociation of the product H4F as the rate-limiting step in the steady-state cycle. The assistance of the cofactor NADPH in promoting product dissociation is an unusual feature, though not limited to DHFR, and follows the rapid loss of NADP+.
Events around the chemical step of reduction/oxidation were monitored by directly observing the conversion of NADPH to NADP+. The kinetics are again biphasic owing to the rapidity of the hydride transfer process; that the rapid phase is associated with the chemical step is verified by the observation of a kinetic deuterium isotope effect of 3 when the transferring hydrogen of the NADPH is replaced with deuterium. This step shows a pH dependence with a pKa of 6.5 that implicates the Asp125 (27 in E. coli) in the proton transfer events required to complete the reduction.
FIGURE 5 The mechanism of amide hydrolysis by a-chymotrypsin.
FIGURE 6 The kinetic scheme for conversion of H2F to H4F by DHFR, including the rate constants for each step at 25°C. In this scheme, NH represents NADPH and N+ represents NADP+.
Measurement of this step in the reverse direction (i.e., for DHFR ■ NADP+ ■ H4F) coupled with determination of the overall equilibrium constant permitted construction of Fig. 6.
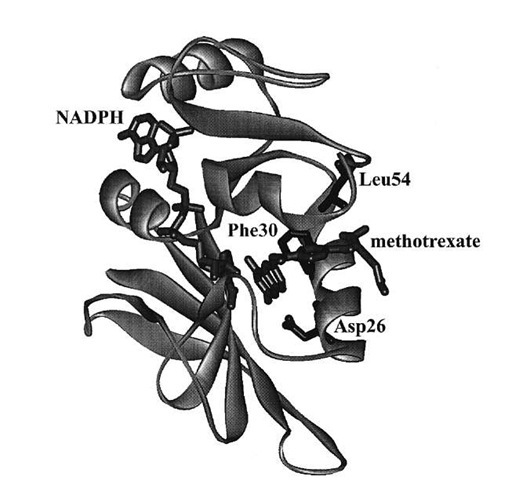
The kinetic scheme served as the basis for the explanation of the contribution of various elements of the protein to its function. Site-specific mutagenesis is a technique in which one or more amino acids are replaced by other amino acids through alteration of the gene encoding the enzyme. For the mutant proteins, the same kinetic scheme was reconstructed to calculate the free energy differences arising from changes in the kinetic steps caused by the mutations. Replacing the hydrophobic residues such as Phe30 and Leu54 (Fig. 7) singly or pairwise with other amino acids revealed that the cumulative effect of two mutations was generally nonadditive in terms of the free energy associated with individual steps in Fig. 6, consistent with long-range interactions across the enzyme active site mediated by bound substrate and cofactor. The nonadditivity differed for each step in Fig. 6, which implicated differing conformations of the protein as arising throughout the catalytic cycle.
Of particular interest was the discovery that changes in the amino acid sequence at loci outside the active site also strongly influence (by a factor of > 102) the rate of the chemical step. In combination with dynamic NMR measurements and molecular mechanics calculations, this observation has been attributed to the importance for catalysis of long-range motions that occur across the entire DHFR protein molecule. The protein fold through its complex vibrational modes apparently may couple some set of motions to a promotional vibration that fosters passage of the reactive ternary complex over the activation barrier.
FIGURE 7 Crystal structure of DHFR from Lactobacillus casei with methotrexate (a strong inhibitor) and NADPH bound. Amino acid residues discussed in the text are labeled.
C. Phosphate Transfer
Enzymes that catalyze the transfer of a phosphoryl moiety between two substrates have provided excellent examples of the use of isotopes in kinetic and stereochemical studies. The enzyme hexokinase, which promotes the conversion of glucose plus ATP to glucose-6-phosphate and ADP has been the subject of kinetic studies that suggested an ordered kinetic sequence with glucose being the first substrate to add and glucose-6-P the last product to be released. Specific information on the identity of rate-limiting steps and the steady-state levels of reaction intermediates was obtained by isotope trapping studies. In its simplest form, enzyme and isotopically labeled substrate (S*) are incubated (the pulse) and rapidly diluted into excess un-labeled substrate (the chase), and allowed to react for a chosen time. Then the reaction is stopped by a quenching reagent that jumps the pH or denatures the enzyme. From the amount of E ■ S* converted to product versus that lost to dissociation (replacement by S gives nonlabeled product) the dissociation rate of S* from E and other ES complexes can be calculated.
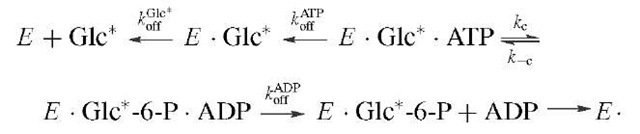
This method has been used in the study of the partitioning of ES complexes in the steady state. In the case of hexokinase, the question was the partitioning of the functional E ■ glucose ■ ATP complex between product formation and substrate release. For glucose the relevant scheme is tainting an 18O label in the j3y bridging oxygen provides the necessary probe for finding this intermediate by means of the process below. In these experiments, isotopes are used as labels so that the fate of a particular atom may be followed throughout the course of the reaction.
In this case the reaction is allowed to reach steady-state turnover, and the solution is either stopped by quench or chased by addition of excess unlabeled substrate followed by a delay sufficient for several turnovers then addition of quench. The presence of a difference in the level of the labeled product obtained by the two procedures represents the concentration of EGlc ATP* complex in the steady state, which is approximately 50% of ET, the total enzyme concentration. The observed steady-state andpre-transient rates are consistent with steps kc and k_c being at equilibrium relative to koAfDf P, which is typical for many phosphotransfer enzymes in which the chemical steps are generally not rate limiting. Additional information can be obtained by using the label in the second substrate (i.e., [y-32P]ATP) and following a similar protocol, which thereby allows calculation of the dissociation rate of ATP from E Glc ATP. In this case E Glc ATP* is approximately 25% of Et, which requires that kAfP compete with the dissociation of ADP (kAfP) from E Glc-6-P ADP. In this manner the individual rate constants for hexoki-nase were largely determined and the order of substrate association was verified.
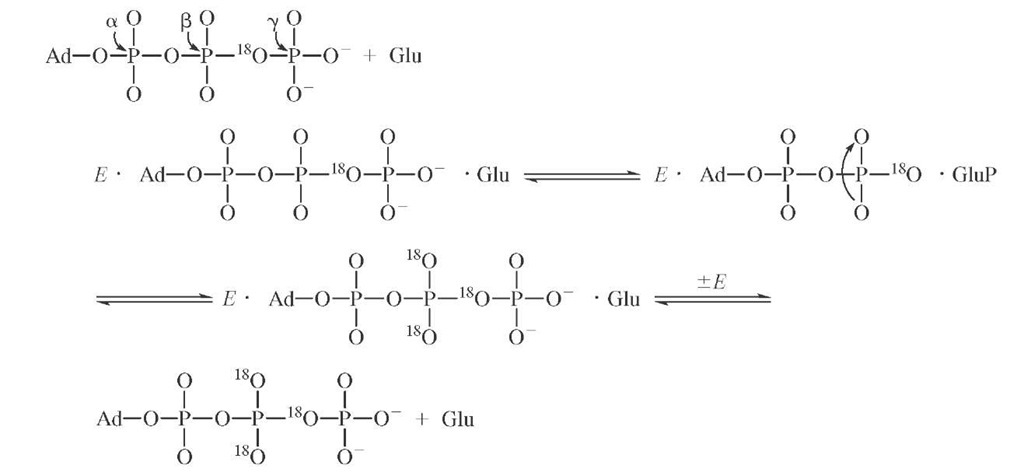
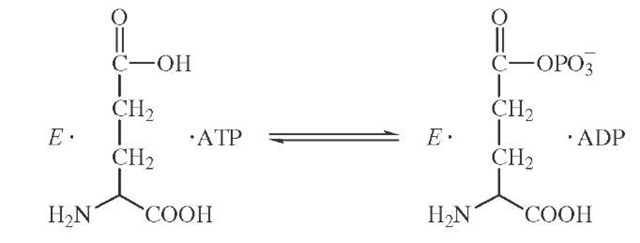
Isotopic labeling has also been cleverly used to demonstrate the existence of enzyme-bound intermediates that do not readily dissociate into solution. The enzyme glu-tamine synthetase catalyzes the formation of glutamine from ATP and ammonia possibly through a tightly bound glutamyl phosphate intermediate.
If the formation of glutamyl phosphate were reversible and occurred in the absence of ammonia, then the presence of a symmetric torsion motion at the cleavage site might be used to detect an isotopic exchange brought about by glutamyl phosphate formation. The synthesis of y-18O4P-ATP con-
The appearance of 18O in the nonbridging oxygens of the j -phosphate can be measured by mass spectrometric and NMR methods. The extent of equilibration is partially inhibited by the presence of ammonia as required if glu-tamyl phosphate is a reaction intermediate.
Isotopic labeling studies of phosphotranferase reactions culminated in the synthesis of ATP chiral at the Y -phosphorus. Chirality was achieved by the synthesis of [Y-16O,17O,18O]ATP of one configuration, and the analysis of its chirality was achieved by stereochemically controlled transfer of the y -phosphoryl moiety to (S)-propane-1,2-diol where the absolute configuration was determined by a chemical/mass spectrometric sequence. The observation of inversion of configuration has been accepted as evidence of an "in-line" displacement mechanism at phosphorus by the two bound substrates; the observation of retention of configuration was used to implicate the existence of a phosphoryl enzyme intermediate in the phosphoryl transfer process. For hexokinase, our case study, the finding is one of inversion, consistent with a direct transfer mechanism.