Agriculture Reference
In-Depth Information
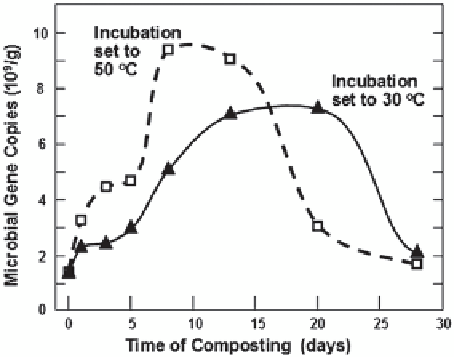
Fig. 3.2
Numbers of cop-
ies of acetomycetes genes
detected in compost mixtures
vs. time, depending on the
temperature of incubation.
(Data replotted from Xiao
et al.
2011
)
higher microbial populations earlier. By contrast, the lower temperature resulted in
slower increases in the number of genes detected, and about six more days had passed
before a major decline in gene count was observed.
3.2.2.1
Quantification of Biota
An ongoing need to quantify populations of bacterial or fungal types in a given
compost mixture has been addressed in various ways in recent studies. Perhaps
the most straightforward approach entails weighing of the microbial biomass pro-
duced under different conditions (Shan et al.
2013
). Reddy et al. (
2011
) and de
Gannes et al. (
2013
) employed pyrosequencing to analyze distributions of microbial
communities. The method makes it possible to distinguish the DNA extracted from
different samples in which microbial communities are present. Li et al. (
2013
) em-
ployed denaturing gradient gel electrophoresis to separate the genes responsible for
enzyme production. As a part of the analysis, the respective genes were first ampli-
fied. Primer molecules having specific DNA sequences were used to amplify the
results of polymerase chain reaction analysis. Wei et al. (
2012
) employed DNA se-
quence detection and were able to document a shift from bacteria-dominated com-
munities to fungi-dominated communities during maturation of compost. Huang
et al. (
2010
) compared the levels of different quinones as a means of judging which
micro-organisms were dominant during different phases of composting. The au-
thors found that certain quinones were indicative of certain fungi.
Another practical way to estimate the relative abundance of different micro-
organisms is by comparing the activities of different identifiable enzymes present
in compost as a function of time (Adams and Umapathay
2011
; Amira et al.
2011
;
Eida et al.
2012
; Gladden et al.
2012
). Adams and Umapathay (
2011
) showed that
extracellular enzyme analysis can be combined with physiological profiling of the
communities of microbes.