Information Technology Reference
In-Depth Information
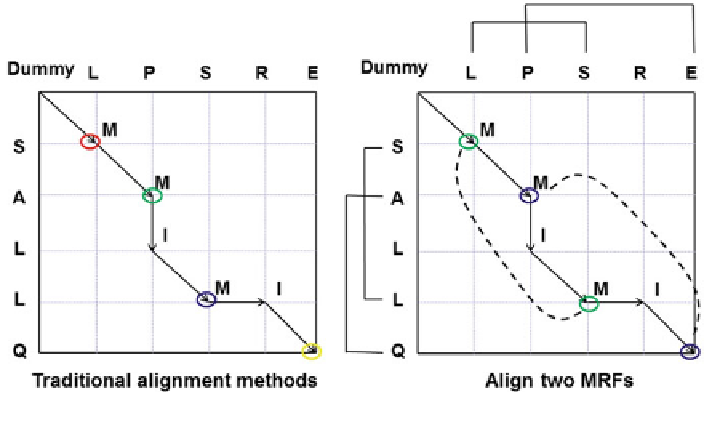
Fig. 2.3 Traditional alignment methods (left) and our MRFalign method (right)
be proved that when edge alignment potential is considered and gaps are allowed
the MRF-MRF alignment problem is NP hard [
10
]. In this work, we will describe
an ADMM [
11
] algorithm to quickly
find a suboptimal alignment of two MRFs.
Although suboptimal, empirically the resultant alignment has very good accuracy.
2.4 Node Alignment Potential of Markov Random Fields
Given an alignment path, its node alignment potential is the accumulative potential
of all the vertices along the path. In particular, the node alignment potential is a
function of two MRF nodes and measures the similarity of two aligned positions
using local protein features. We use a Conditional Neural Fields (CNF) [
12
] method
to develop such a node alignment potential, using a procedure very similar to what
is described in the protein threading paper [
13
,
14
]. Brie
y speaking, we estimate
the occurring probability of an alignment A between two proteins T and S as
follows.
fl
e
P
ð
i
;
j
;
u
Þ2
A
E
u
T
i
;
S
j
ð
=
P
ð
A
j
T
;
S
Þ¼
ZT
ð
;
S
Þ
ð
2
:
4
Þ
where ZT
is a normalization factor summarizing all the possible alignments
between T and S, and E
u
T
i
;
ð
;
S
Þ
S
j
is a neural network with one hidden layer that
calculates the log-likelihood of a vertex
ð
i
;
j
;
u
Þ
in the alignment path, where i is a
node in T, j a node in S, and u a state. When u is a match state, E
u
takes as input the
sequence pro
le context of two nodes i and j, denoted as Ti
i
and S
j
, respectively,
Search WWH ::

Custom Search