Biomedical Engineering Reference
In-Depth Information
GABA
Cl
-
(A)
(B)
ρ
γ
ρ
α
ρ
ρ
β
β
ρ
α
GABA
A
subunits
α1-6, β1-3, γ1-3, δ, ε, π, θ
GABA
C
subunits
ρ1-3
(C)
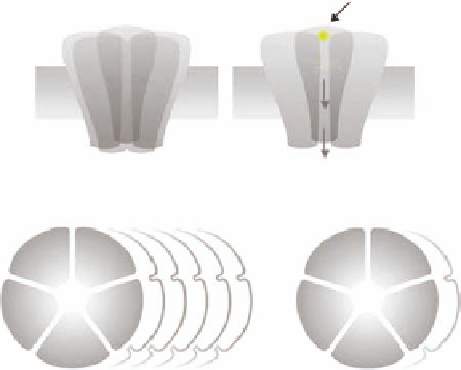
FIGURE 15.5
Schematic illustrations of (A) the pentameric structure of the ionotropic GABA receptors,
(B) with indication of the GABA-binding site and the chloride ion channel, (C) and the multiplicity of ionotropic
GABA
A
and GABA
C
receptors.
are heteromeric complexes, and although a wide range of different receptor combinations exists
in
vivo
,
α
1
β
2
γ
2
combination is the predominant physiological GABA
A
receptor subtype. In contrast,
the GABA
C
receptors are homomeric assemblies of i ve identical
ρ
subunits or pseudoheteromeric
complexes comprising different
subunits.
The binding site for GABA and ligands for the orthosteric binding has been shown to be located
at the interface of the
ρ
subunits in the GABA
A
receptor complex, whereas the binding site
for BZD is located at the interface of the
α
and
β
subunits.
There is still no 3D-structure available for the ionotropic GABA receptors. Thus, the understand-
ing of the molecular architecture of the orthosteric-binding site in the ionotropic GABA receptors
has to a large extent been based on the publication of crystals structures of acetylcholine-binding
proteins (AChBP) from snails. These proteins display low but still signii cant amino acid sequence
homologies with the amino-terminal domains of all ligand-gated ion channels within the Cys-loop
family, including the ionotropic GABA receptors, and this homology has been exploited for the con-
struction of homology models of this region in both GABA
A
and GABA
C
receptors. Such homology
models offer an insight into the identities of the residues lining the binding pockets in the respective
receptors. However, given the low level of sequence identity (~18%) between the AChBP and the
ionotropic GABA receptors, it is not straightforward to use these models for the prediction of ligand
afi nity or structure-activity studies.
α
and
γ
15.5.2 I
ONOTROPIC
GABA R
ECEPTOR
L
IGANDS
The GABA-binding site has very distinct and specii c structural requirements for recognition and
activation. Thus, only a few different classes of structures have been reported. Within the series
of compounds showing agonist activity at the GABA
A
receptor site are the selective agonists mus-
cimol (
15.22
) and THIP (
15.23
), which have been used for the pharmacological characterization
of the GABA
A
receptors. BMC (
15.24
) and SR95531 (
15.25
) are the classical GABA
A
receptor
antagonists.
In the absence of a 3D-receptor structure, the relationship between the ligand structure and
the binding/activity at the GABA
A
receptor has been extensively studied. On the basis of a