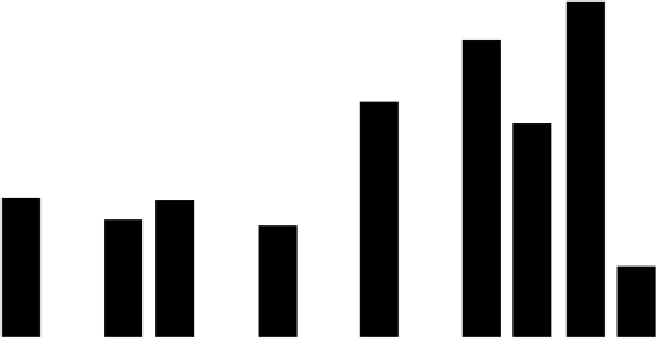Biomedical Engineering Reference
In-Depth Information
demonstrated that these values correlated linearly with the logarithm of the measured
chromosomal loss rates (170). This type of correlation between a simple calculable DNA
physical property for any altered DNA sequence in a DNA-protein complex and the meas-
ured functional property of the altered sequence complex was a novel observation. Also,
for each CDEI and CDEIII binding site position, plots of the change in PD values
vs. log(chromosomal loss rates) for the three possible mutants and the wild-type sequence
were made and their slopes determined. As Figure 1.50 illustrates, these slope magnitudes
for each CDEI and CDEIII site position were found to be symmetric about the center of
symmetry of these binding sites. That is, position pairs 3 and 8, 4 and 7, and 5 and 6 pos-
sess similar slopes in CDEI; and position pairs 11 and 17, 12 and 16, and 13 and 15 possess
similar slopes in CDEIII (170). This symmetric site behavior agrees with the known rota-
tional symmetry of the conserved CDEI and CDEIII binding sites and provides stronger
validation that our simple PD-scale-based computational approach is revealing important
quantitative information about DNA-protein complexes.
These results support the validity of our approach using the PD scale parameters to
begin to understand the energetic basis of DNA sequences interacting with their recogni-
tion proteins. One advantage this approach provides is that DNA sequence-based physi-
cal properties are far easier to calculate than protein physical properties. This results from
the fact that all DNA as it occurs inside cells has a regular repeating secondary structure
close to the B secondary structure proposed originally by Watson and Crick (171). As a
consequence, a number of different dinucleotide- and trinucleotide-based DNA physical
property data sets (PD, bendability, and Position Preference) have been calculated from
experimental data (172). Using these data sets, the local physical properties for any DNA
sequence can be calculated. No such equivalent simple computational rules exist for pro-
teins, which lack the overall energetic and structural uniformity that DNA possesses. The
computational approach we have described has important implications for the use of
DNA-protein complexes in biosensors. Stability and property differences for DNA
0.6
V = Centers of symmetry
V
0.5
0.4
0.3
V
0.2
0.1
0
1
2
3
4
5
6
7
8
11
12
13
14
15
16
1
7
CDE I
CDE III
FIGURE 1.50
Histogram plot of the slopes determined from the experimental log
e
(chromosomal loss rates) vs.
PD plots cal-
culated for each binding site position of CDEI and CDEIII. The site position numbers are indicated along with
the center of symmetry (V symbol) for each binding site. From Henemuth, B., Marx, K.A. (2006). Protein
Deformability Properties of Single Base Mutants of Yeast DNA Kinetochore Protein Binding Sites Correlate with
Measured Chromosomal Loss Rates for the Mutants in Yeast Cells.
BMC Mol. Biol.
7:12-20. With permission.