Information Technology Reference
In-Depth Information
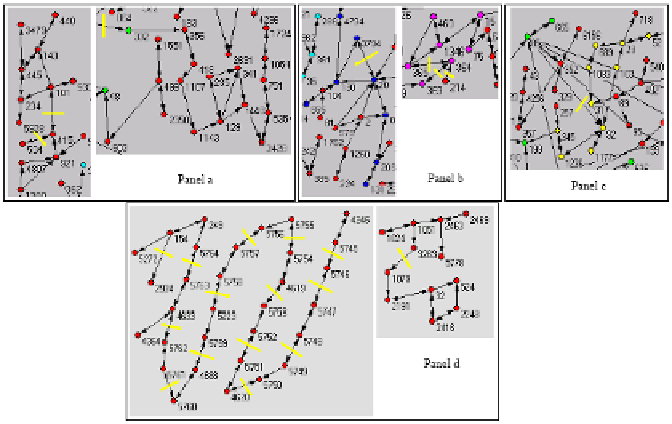
Fig. 8.4. Examples reporting the main effects of enzyme inhibitions. Panel a: isolating and
separating a pathway from the rest of the network. Panel b: interrupting the full connectivity in a
strongly connected component. Panel c: creating a new smaller SCC from another one. Panel d:
disrupting a cluster of nodes in the IS compartment.
The analysis of the metabolic network wiring after the deletions
corresponding to the knocking out of an essential enzyme, allowed us to discover
that all of the enzymes corresponding to lethal mutations, when deleted, prevent
the connections between the separated nodes. No other path is available to restore
the broken arc and the involved metabolites (nodes) are no more connected by
alternative pathways. The strict relation between lack of alternative pathways
reaching a given metabolite and enzyme essentiality implies that the crucial
elements of the network correspond to periphery edges (more central locations
being more easily reachable by alternative paths along the graph): this is in line
with the recently discovered importance of the so called non-hub connectors
(elements linking different modules of the network).
The a-la-Barabasi approach to the study of scale-free networks is based on the
effects produced by the elimination of nodes. In this perspective, essential nodes
in the protein interaction network are often hubs (nodes at the center of the
connection pattern). By contrast, the relevance of periphery in metabolic
networks is revealed by considering the elimination of edges which corresponds
to the elimination of the associated enzyme(s).
The factors influencing the relevance of the elimination of a node or an edge
on the overall network are exactly the opposite. In fact, the entity of the
“damage” produced by the elimination of a node is proportional to the number of