Information Technology Reference
In-Depth Information
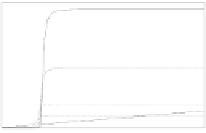
Inverter: Modify Repressor Affinity
30
baseline inverter
repressor 10x weaker
repressor 100x weaker
repressor 1000x weaker
x=y
25
20
15
10
5
0
0
5
10
15
20
25
30
psi(A) = input signal
Figure 4.20
Reducing the binding affinity of the repressor to the operator shifts the
transfer function of an inverter outward.
therefore less mRNA output (
ψ
Z
). BioSPICE simulations (Figure 4.21) show
the overall inward shifts of the transfer curve of the
cI
/P(R) inverter that result
from weakening the P(R) promoter. In these simulations, the original value of
k
xscribe
was scaled down by factors of 2, 5, and 10.
MODIFYING THE STRENGTH OF THE RIBOSOMAL BINDING SITE (RBS) Modifica-
tions to the RBS alter the affinity of rRNA to the mRNA. Figure 4.19c illustrates
the shift in
L
, the translation stage mapping between
φ
A
, resulting from
a reduction in rRNA affinity. For any given level of the input mRNA,
ψ
A
and
ψ
A
, there
Modify promoter affinity for inverter (inset: two inverters)
baseline inverter
promoter 2x weaker
promoter 5x weaker
promoter 10x weaker
x=y
30
25
30
20
25
20
15
15
10
5
10
0
0
0.5
1
1.5
2
2.5
3
3.5
4
5
0
0
0.5
1
1.5
2
2.5
3
3.5
4
psi(A) = input signal
Figure 4.21
Reducing the binding affinity of the RNA polymerase shifts the transfer
functions of the
cI
/P(R) inverter inward. Inset shows the effects on the transfer func-
tions of two inverters in series. The diagonal lines correspond to input equals output.