Information Technology Reference
In-Depth Information
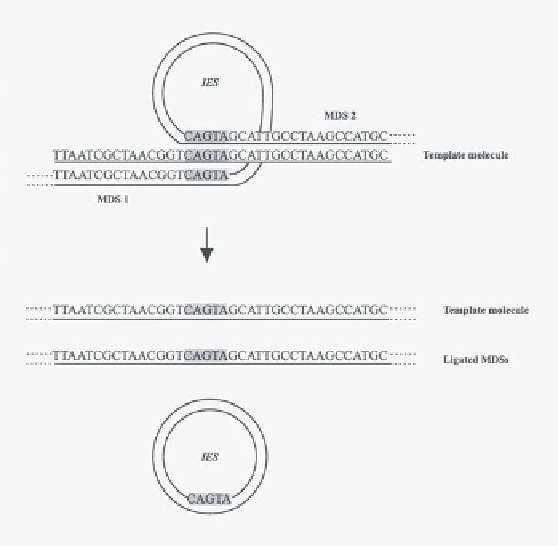
Figure 9.29
General view of the template-guided model of gene assembly in sti-
chotrichs. A DNA molecule from the old macronucleus aligns macronuclear destined
segments (MDSs) in the orthodox order by homologous pairing with the MDSs. Such
alignment brings the correct pointers (CAGTA) into position for recombination in
an ld excision operation. The nucleotide sequences are shown for the 5'
3' chain
of DNA double helices. The complementary chains are shown as lines. The excised
internal eliminated segment (IES) is shown as a circle, but the excised IES could be
linear, depending on how the recombination is postulated to occur.
→
We propose a template-guided model of gene assembly to assure alignment
of correct copies of pointers. The model is based on work with
Paramecium
in the laboratories of Meyer [1] and Forney [4] that shows that the old macro-
nucleus can guide IES excision in a new, developing macronucleus. The essence
of the template model for gene assembly in stichotrichs, which is described
in detail in Prescott, Ehrenfeucht, and Rozenberg (submitted), is shown in
Figure 9.29.
In the model, copies of DNA molecules from the old macronucleus enter the
developing macronucleus and align by homologous pairing to MDS sequences
in the DNA of the polytene chromosomes. In Figure 9.29 alignment of an old
macronuclear sequence with two numerically consecutive MDSs in the DNA in
polytene chromosomes aligns the pointers in those MDSs, producing a looping
out of the IES. Recombination between members of the pointer pair excises