Biology Reference
In-Depth Information
Subgroup 4-
related sequences
42
Pg
MYB16
42
99
Zm
MYB31
Subgroup 4
Monocotyledoneous
subclade
Subgroup 4 gymnosperms
monophyletic clade (Sg4C)
Subgroup 4
angiosperms clade
0.1
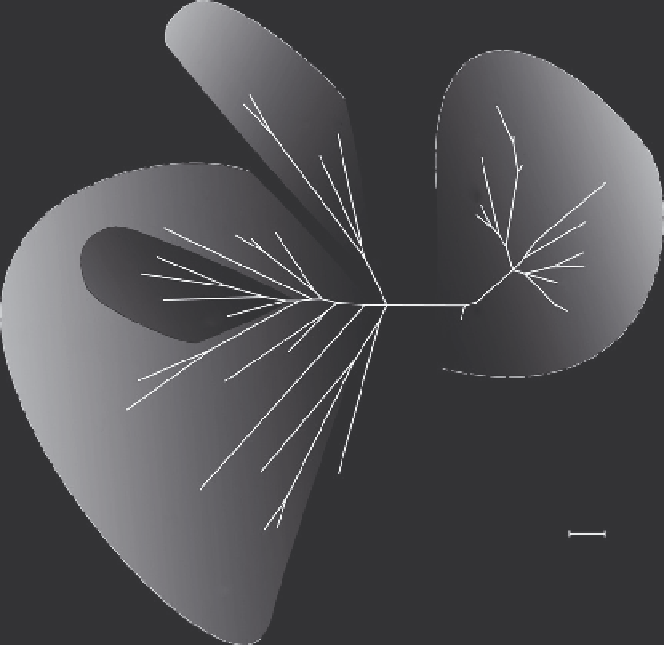
Fig. 2. Phylogenetic tree was constructed using the R2R3-MYB genes from the
subgroup 4, using the sequences from Arabidopsis (
Dubos et al., 2010; Stracke et al.,
2001
; including the subgroup 4-related sequences AtMYB6 and AtMYB8), the
sequences from Pinus taeda and Picea glauca described in
Bedon et al. (2010)
,
the predicted sequences from the Populus trichocarpa genome putatively belonging
to this subgroup (
Tuskan et al., 2006
), and the landmark genes belonging to subgroup
4 described in
Table 1
. Since many sequences from P. taeda and P. glauca lack the first
35 amino acids, 35 amino acids in the N-terminal region, (starting with the second
tryptophan of the R2R3-MYB domain) were deleted from all of the sequences used
for the tree. The methodology used to construct the tree is the same as described in the
legend of
Fig. 1
. All the gymnosperm sequences, except PtMYB22, are grouped
together in a big monophyletic subclade named Sg4C, as described by
Bedon et al.
(2010)
, although PgMYB13 and specially PtMYB13 are more distant from the other
gymnosperm genes within this subclade. For the angiosperm sequences, a specific
clade contains all the monocot sequences, including wheat, maize and switchgrass.
Moreover, most of the poplar sequences are grouped by two, suggesting that this
could be a result of the salicoid-specific whole genome duplication event in the
Populus lineage (
Wilkins et al., 2009
).