Information Technology Reference
In-Depth Information
0.7
35
0.6
30
0.5
25
0.4
20
0.3
15
0.2
10
0.1
5
0.0
5
10
15
20
25
30
35
Locus
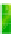
Fig. 5.
Heatmap of activation probabilities for main effect and epistatic effects for the
Arabidopsis
thaliana
Bay-0
×
Shahdara at the 38 loci. Activation probabilities along the diagonal correspond
to main effects and off diagonal correspond to epistatic effects.
all interaction terms
jk
in model
i
;for
σ
2
,
λ
=1
and
ν
=1
are used. Note that when
ν
=1
and
lambda
=1
the
Inv
χ
ν,λ
has infinite mean and variance. Hence, should
be relatively uninformative. For each model
M
c
where multicollinearity does not occur,
the prior probability
P
(
M
c
)
is chosen uniformly across this space. Thus, a priori, no
model is preferred over another.
Using the restriction
r
=10
, 25 chains of 11,000 were run with a burn in of 1,000
samples using overdispersed starting models resulting in 250,000 samples.
The number of visits to model
M
c
was recorded and the probability of model given
the data
P
(
M
c
|
−
D
)
was estimated as the number of visits to model
M
c
divided by the
length of the chain. The probabilities appeared to converge after 15 chains, indicating
convergence. Using these probabilities the activation probabilities
P
(
β
j
=0
|
D
)
and
P
(
β
j
k
D
)
are computed for each main and epistatic effect, respectively. Figure 5
shows a heat map of the epistatic probabilities. Notice that the highest activation prob-
ability locus on the heat map is at locus 18 (ATHCHIB2) and no epistatic, off diagonal,
effects have high activation probability.
The activation probabilities for the 3 highest loci, ATHCHIB2, MSAT5-9 and MSAT5-
22 are as follows:
P
(
ATHCHIB
2
=0
|
|
D
)=0
.
741
,
P
(
MSAT
5
−
9
|
D
)=0
.
481
and
P
(
MSAT
5
D
)=0
.
445
. This suggests that the following epistatic effects should
be considered:
ATHCHIB
2
−
22
|
×
MSAT
5
−
9
,
ATHCHIB
2
×
MSAT
5
−
22
and
MSAT
5
22
. The marginal epistatic activation probabilities and the
conditional epistatic activation probabilities of these interactions are shown in Table
1. Previous studies have shown ATHCHIB2 to be a locus associated with cotelydon
opening [4]. Hence the results agree with biological expectations. In order to more ac-
curately locate the locus associated with cotelydon opening a dense map of genes near
ATHCHIB2 should be undertaken.
−
9
×
MSAT
5
−