Biology Reference
In-Depth Information
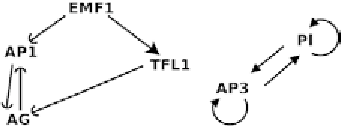
Fig. 8. The simplified
A. thaliana
flower morphogenesis gene regulatory network.
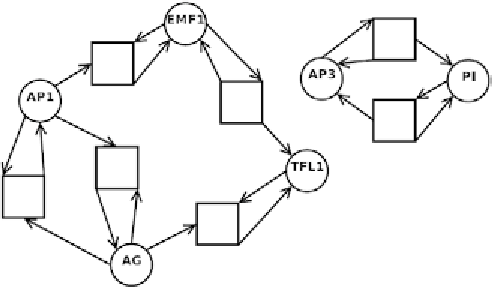
Fig. 9. Petri net corresponding to the gene regulatory network in Fig. 8.
An advantage of the straightforward approach is that no preprocessing of the network is necessary.
However, it produces many false-positives and many genuine states are not identified. For instance the
states [0001000000] (equivalent to activity
A
that is observed in
sepal
cells) and [0001000110] (equivalent
to activities
A
and
B
that are observed in
petal
cells). This implies that substantial post-processing is
necessary.
The refined approach
The refined approach overcomes the weakness of the straightforward approach, although at the cost of
some preprocessing. In addition the resulting network is simpler, and therefore the
Presburger arithmetic
formulae are much shorter. The idea is to remove the temporary genes from the analyzed network. So
the simplified network consist only of genes
EMF1
,
TFL1
,
AP1
,
AG
,
AP3
and
PI
, as shown in Fig. 8.
Figure 9 shows the corresponding Petri net.
The formula below is an instance of Eq. (3), for the subnetwork consisting of genes
PI
and
AP3
. The
variable
t
AP I
denotes the activation of
PI
by
AP3
and variable
t
IPA
denotes the activation of
AP3
by
PI
.
⎛
⎝
∀
⎛
⎝
(
t
AP I
1
∧
(
PI
1
∧
AP
3
1)
∧
t
IPA
1)
∧
t
API
,t
IPA
place bounds
possible transitions
⎛
⎝
non conflicting (
P
I
,AP
3
,t
AP I
,t
IPA
)
⎛
⎝
∀
t
API
,t
IPA
t
AP I
1
∧
t
IPA
1
∧
∧
non
−
conflicting set
primed
variables denote the superset