Biology Reference
In-Depth Information
(a)
(b)
S
UB1
B
S
UB1
C
SUB1A
S
UB
1B
SU
B1A
-
2
S
UB1
C
SU
B1A
-
1
S
UB
1B
S
UB1
C
SUB1A-1
SUB1A-2
∗
(c)
(d)
+Sub1
Submergence
Ethylene
ROS-scavenging
Recovery growth
-Sub1
SUB1A-1
Cell elongation
CHO consumption
Chl degradation
SLR1, SLRL1
Leaf elongation
CHO consumption
Chl degradation
Cell death
GA response
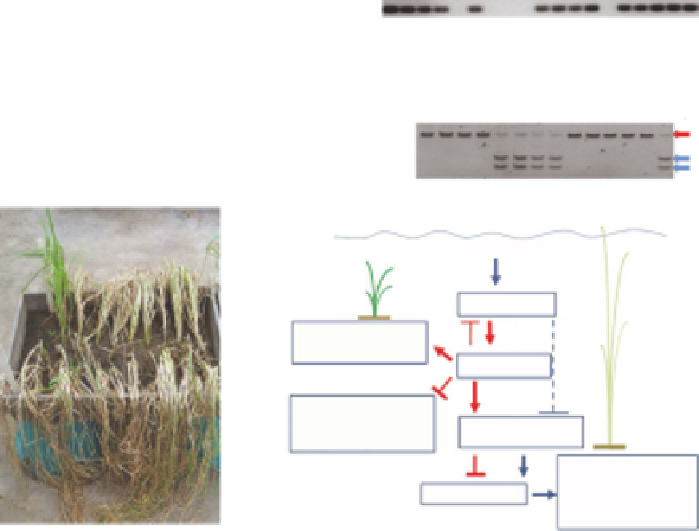
Fig. 2.3.
SUB1
-specific markers and the molecular basis of submergence tolerance.
The
SUB1
locus contains either
two (
SUB1B
,
SUB1C
)orthree(
SUB1A
,
SUB1B
,
SUB1C
) ERF genes (
a
). Two different indica-/aus-specific
SUB1A
alleles
are known, of which the
SUB1A-1
allele is present in submergence-tolerant varieties. Molecular markers distinguish
between varieties with and without
SUB1A
(
b
, top panel) and between the
SUB1A-1
and
SUB1A-2
allele (
b
, bottom
panel). The visible effect of the
SUB1
locus is the suppression of growth during submergence and plant recovery within
about 2 weeks after de-submergence (
c
). Under submergence, ethylene induces the gibberellic acid (GA)-dependent
escape response, which is suppressed by
SUB1A-1
-mediated maintenance of GA repression via the GA repressor proteins
SLR1 and SLRL1 (
d
). ROS, reactive oxygen species; CHO, carbohydrate, CHL, chlorophyll, GA, gibberellic acid, SLR1,
SLENDER RICE 1, SLRL1, SLENDER RICE LIKE 1. For a color version of this figure, please refer to the color plate.
SUB1A-1
is also the allele present in the
SUB1
donor variety FR13A and FR13A-derived lines
that are now being used in Sub1 breeding pro-
grams (see above). The molecular markers used
for Sub1 breeding enable breeders to distinguish
between plants with and without the
SUB1A
gene, and to discriminate between the alleles of
SUB1A
(Figure 2.3b).
It has recently been suggested that the
SUB1A-2
allele can also confer tolerance of sub-
mergence since high expression of the
SUB1A-
2
allele was observed in some varieties with
intermediate tolerance (Singh et al. 2010; Sep-
tiningsih et al. 2012). However, the most tol-
erant
SUB1A-2
variety (James Wee) had a
submergence-survival rate of 44%, which is sig-
nificantly lower than
SUB1A-1
varieties (51-
65%) (Singh et al. 2010). It remains to be
shown whether the higher tolerance mediated
by
SUB1A-1
compared with
SUB1A-
2 is related
to the level of transcription or post-translational
modification, for example, phosphorylation at
the putative MAP-kinase target site that is spe-
cific to the
SUB1A-1
allele (Xu et al. 2006).
The main effect of
SUB1A-1
is the sup-
pression of the “escape” response, which, in

























Search WWH ::

Custom Search