Biology Reference
In-Depth Information
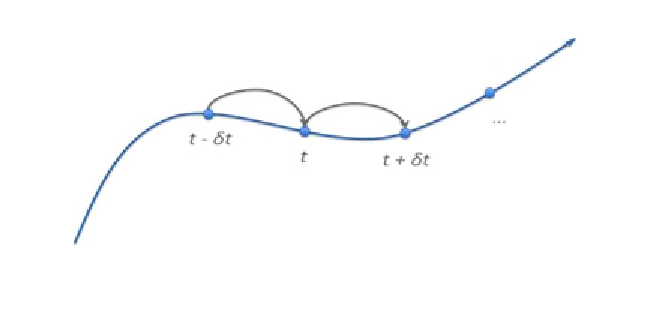
Fig. 2.
Graphical representation of the stepwise time solution obtained by the
Verlet algorithm.
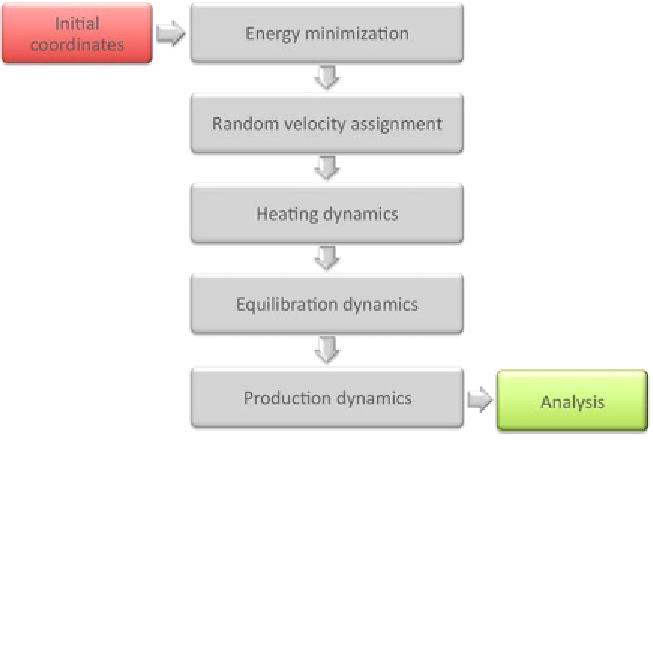
Fig. 3.
Main steps involved in the MD simulation of a given structure.
obtained from an experimental structure (X-ray or NMR) or a theoreti-
cal model (homology modeling or
ab initio
folding) are first minimized
to prevent large unphysical motions at the start of the dynamics result-
ing from potential steric clashes. Initial velocities are assigned randomly