Chemistry Reference
In-Depth Information
f
(
x
)
Local search
Phenotypes
Lamarckian
Inverse
Mapping
Mutation
Mapping
Genotypes
Child
Parent
Child
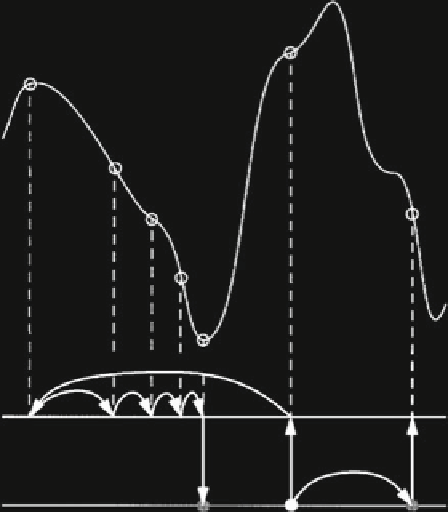
Fig. 3 An illustration of the genetic algorithm approach, where the states of the ligand (transla-
tion, orientation, and conformation relative to the protein) are interpreted as the ligand genotype
and the atomic coordinates represent the phenotype. A plot of the change in the fitness function
(
f(x)
) as the ligand population is allowed to mutate, crossover, and migrate. The genetic evolution
of the ligand effectively samples conformational space where the best conformer is identified by
a minimum in the fitness function (Reprinted with permission from [
179
], copyright 1998 by
John Wiley and Sons)
utilize a tabu list to remember previous ligand states. A pose is immediately
rejected if it is close to a prior conformation. The tabu list encourages the search
to progress to unexplored regions of conformational space.
3.2 Scoring
While docking algorithms are generally efficient at generating the correct ligand
pose, it is important for the docking program to actually select the correct ligand
conformation from an ensemble of similar conformers. In essence, the scoring
function should be able to distinguish between the true or optimal binding confor-
mation and all the other poses. The scoring function is also used to rank the relative
binding affinities for each compound in the library. Ideally, the scoring function
should be able to calculate the free energy (
G
binding
) of the protein-ligand binding
interaction, which is directly related to the
K
D
:
D