Information Technology Reference
In-Depth Information
1
0.8
0.6
0.4
HKM
HKM+EI
0.2
0
0
0.2
0.4
0.6
0.8
1
False positive rate
Fig. 2.
ROC analysis to show the effect of evolutionary information on random forest classifi-
ers constructed using the PDNA-62 dataset with the ASA-based definition of DNA-binding
residues. HKM represents the classifier trained with the three biochemical features (
H
,
K
and
M
), and HKM+EI indicates the classifier using two types of evolutionary information (PSSM,
H
m
,
H
d
,
K
m
and
K
d
).
the best classifier trained with sequence identity and entropy achieved 78% overall
accuracy but with only 41% sensitivity and MCC = 0.28. Although a different dataset
was used for classifier construction and evaluation in the previous study [5], the RF
classifier developed in the present study appears to be significantly more accurate than
the Naïve Bayes classifier for DNA-binding site prediction. It is likely that the use of
evolutionary information together with biochemical features for input encoding in this
study but not in the previous study [5] is responsible for the improved classifier per-
formance.
3.3 Classifier Evaluation Using a Separate Test Dataset
The results presented so far have been obtained from fivefold cross-validation ex-
periments on the PDNA-62 dataset. To further evaluate the most accurate RF in Table
2 (also called BindN-RF), we prepared a separate test dataset (PDC25t), which shared
less than 25% sequence identity with the PDNA-62 dataset. The RF classifier was
also compared with two of the previously published classifiers (BindN and DBS-
PSSM). BindN used the SVM classifier constructed using the three biochemical fea-
tures in our previous study [10]. DBS-PSSM (http://www.netasa.org/dbs-pssm/) used
the ANN predictor trained with PSSM and sequence information [6]. These two exist-
ing classifiers were chosen because they were constructed using the same training
dataset (PDNA-62) as in the present study, and used the same distance-based criterion

















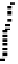






Search WWH ::

Custom Search