Information Technology Reference
In-Depth Information
50
−
150
Box.H
50
−
100
0
Tru.H
T68(15)
40
0
Box.L
−
50
Ear.L
Tru.L
0
T64(30)
50
−
0
Cra.L
Ear.H
0
Fow.H
Hun(62)
20
20
Edn.H
−
50
0
Cap(72)
Beg.L
Cra.H
50
0
50
Edn.L
0
Beg.H
0
−
50
50
Hob(78)
100
−
50
Fow.L
0
−
50
0
100
−
100
100
Spo(44)
50
T95(8)
−
100
40
−
50
(b)
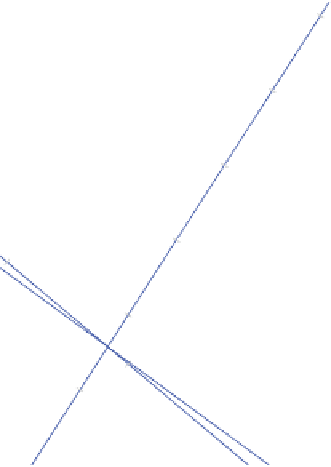
Figure 6.8
(
Continued
)
circle.proj = 5
. Similarly, finding for any given variety the interaction predictions
at all sites is achieved by construct a circle with diameter the line connecting the variety
with the origin. This is shown for variety
Ran
in Figure 6.10 by calling
biadbipl
with arguments
X = t(wheat.data), circle.proj = 2
. The reader can verify that
the predictions obtained from Figures 6.9 and 6.10 are as given in Table 6.6. Note
that the predictions in Table 6.6 are readily obtainable by utilizing the argument
pre-
dictions.sample
when calling
biadbipl
. The reader can verify that this argument
allows connecting lines to be drawn to the predictions in the circles shown in Figures 6.9
and 6.10.
If preferred, the origin of the biplot axes in Figures 6.9 and 6.10 can be interactively
shifted to a more convenient position by setting
select.origin = TRUE
in the call to
biadbipl
. Examples are shown in Figure 6.11.
A biadditive biplot showing the predictions at all sites for the same selected variety
or the predictions for all varieties at any selected site can easily be constructed by
calling
biadbipl
with arguments
X
or
t(X)
together with an appropriate value
given for the argument
predictions.allsamples.onaxis
. This is illustrated in
Figures 6.12 - 6.14. The reader may check the predictions from Figures 6.12 and 6.13 by
calling
biad.predictivities
appropriately. In Figure 6.14 we show how to obtain
predictions (in terms of interactions with main effects added) in a one-dimensional
biadditive biplot.