Biology Reference
In-Depth Information
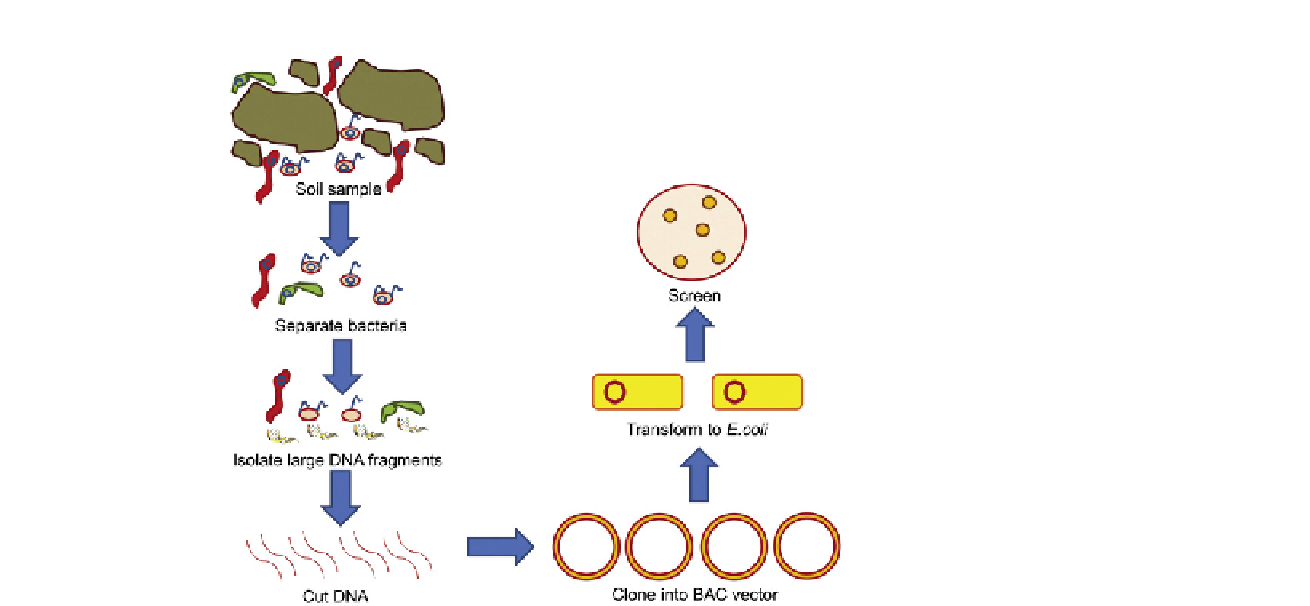
FIGURE 10.1
Process flow of the metagenomic method.
in bacteria are typically clustered together on the genome. Therefore, this method is widely
used to identify new biologically active compounds.
191
There are two major metagenomic screening strategies: expression-dependent (function-
based) and expression-independent (sequence-based). Both have been used to identify
eDNA clones that produce bioactive compounds. In expression-dependent studies, the
production of bioactive compounds is usually linked to the change of phenotypes generated
from the eDNA libraries through a simple high-throughput assay. On the other hand, in
expression-independent studies, the eDNA libraries are examined by screening methods that
use probes containing conserved sequences traditionally associated with secondary
metabolite biosynthesis to identify target clones. Valuable information collected from these
initial high-throughput assays is subsequently tested in model cultivable heterologous
hosts to confer the production of desirable compounds.
84
There are several examples of using the function-based metagenomics library screening
method. One new isocyanide-containing eDNA-derived antibiotic was isolated and
characterized by Brady and coworkers.
85
In this work, eDNA clones in
E. coli
libraries were
screened using top agar overlay assays to identify potential bioactive compounds by looking
for clones generating inhibition zones against test microbes. In addition, a variety of
long-chain
N
-acylated amino acid antibiotics were characterized by the same method.
86
Antibacterially active pigments violacein, indigo,
81,87
and turbomycins
88,89
were also found
by examining pigmented eDNA clones or through randomly selected culture broth
extracts.
90
92
Compared to the function-based metagenomics library screening method, the sequence-based
metagenomics library screening method offers a better way to clone natural product gene
clusters from uncultivable symbionts or gene cluster families from the environment. The
anticancer drug pederin was originally isolated from the beetle
Paederus fuscipes
,
93
while the
biosynthetic gene cluster for pederin was identified to originate from an uncultured symbiotic