Biology Reference
In-Depth Information
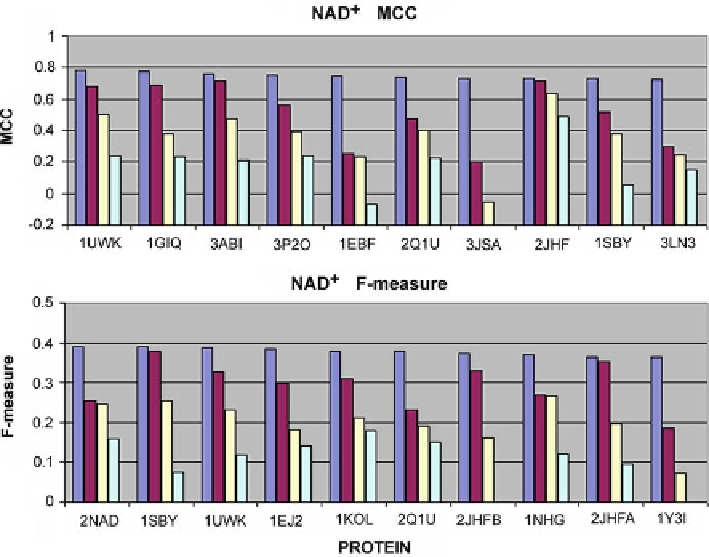
Fig. 4.13
MCC and F-measure scores for NAD
+
-complexing proteins correctly processed by most
of the analyzed tools. Only the top four scores have been taken into account for each program
tools are likely to contain well-ordered pockets whose geometry closely corresponds
to the requirements of a specific ligand. This hypothesis is supported by the high
consistency of results returned by various geometry-based packages. A selection of
best- and worst-case identification scenarios is presented in Figs.
4.15
and
4.16
.
An interesting case is the 2Z6I protein (oxidoreductase E.C. 1.3.1.9 in complex
with the FMN ligand). It was accurately recognized by the FOD model, as well as
by QSiteFinder, CASTp, SuMo and PocketFinder. 2Z6I is a globular molecule with
a deeply embedded ligand which affects the structure of the hydrophobic core and
suggests significant distortion (hydrophobicitiy deficiency) caused by the binding
pocket depth (Fig.
4.17
).
Similarly, accurate FOD results were obtained for the 1N9L protein (electron
transport - putative blue light receptor). These results are presented in Fig.
4.17
.
Again, we are dealing with a globular molecule with a deeply embedded FMN
ligand. Plotting ROC curves for both proteins (and particularly for 2Z6I) reveals
that the area bounded by each curve and the corresponding diagonal is quite large.
Residues responsible for binding ligands are highlighted on
Δ
H
pro fi le plots.
Results for a cutoff threshold of 0.004 are presented in Table
4.6
.
Table
4.6
compares proteins which yielded correct and incorrect results when
applying the ROC curve method (based on the FOD model). The criterion of cor-
rectness is the area bounded by the ROC curve and the corresponding diagonal.
Search WWH ::

Custom Search